This arose out of a thread on social media
Where can I find a practioner well versed on gut microbiome, mold, and possibly gastrology? I’ve been getting worse the last few years with my functional provider that I’m now dealing with low ferritin, oxalates, mold, Sibo, Candida, and malnutrition. This has thyroid and hormones out of whack along with increasing TM.
Response:The harsh reality is many promises, very few deliver. A Colleague is spending many hours a week trying to instruct medical professionals on the gut microbiome… he says that it is painful…
Why is this so?
Looking at studies on the microbiome on PubMed we see an explosion of knowledge. 228,197 studies cited for “microbiome”. The consequences are simple:
- Any medical professional that graduated before 2017 likely has had near-zero training on the microbiome.
- Medical professional tend to use “established cook-book recipes” for treatment. One key reason is medical liability insurance.
- For a microbiome issue like Ulcers, it took almost 50 years for the treatment cookbook to be widely adapted.
- Antibiotics working was reported in the 1950’s
- Barry Marshall and Robin Warren in 1979 identified the bacteria
- In the 1990’s the FDA approved the use of antibiotics.
- For a microbiome issue like Ulcers, it took almost 50 years for the treatment cookbook to be widely adapted.
My father suffered (nearly dying) from a bleeding ulcer in the 1960’s. That reality contributed to my not wanting to wait until treatments are approved by medical insurance companies.
Solutions
Conventional
There are a small number of people with the appearance of skills. I have a list here. The people have not been evaluated (buyer beware), but they are at least interested. I know that Kristina Mitts actively uses microbiome prescription and is also a writer for BiomeSight ,example of one of her posts.
Self-Serve
I suffered from ME/CFS (Chronic Fatigue Syndrome) multiple times and have a lot of contacts that still have it. Most of those people cannot afford to go conventional. The same people also suffer from brain challenges, so I started up a blog site for them: CFS Remission. That site spawned my Microbiome Prescription site.
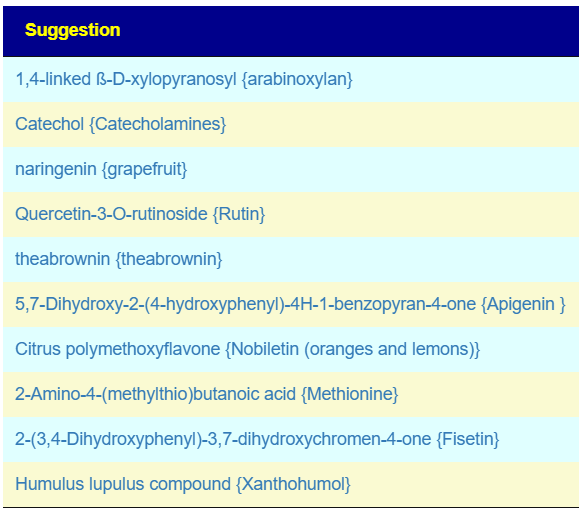
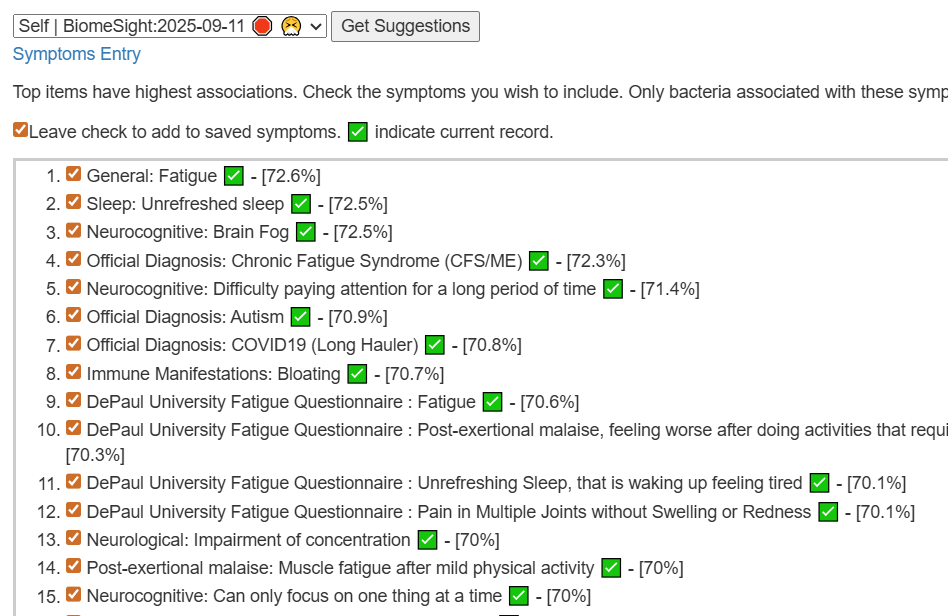
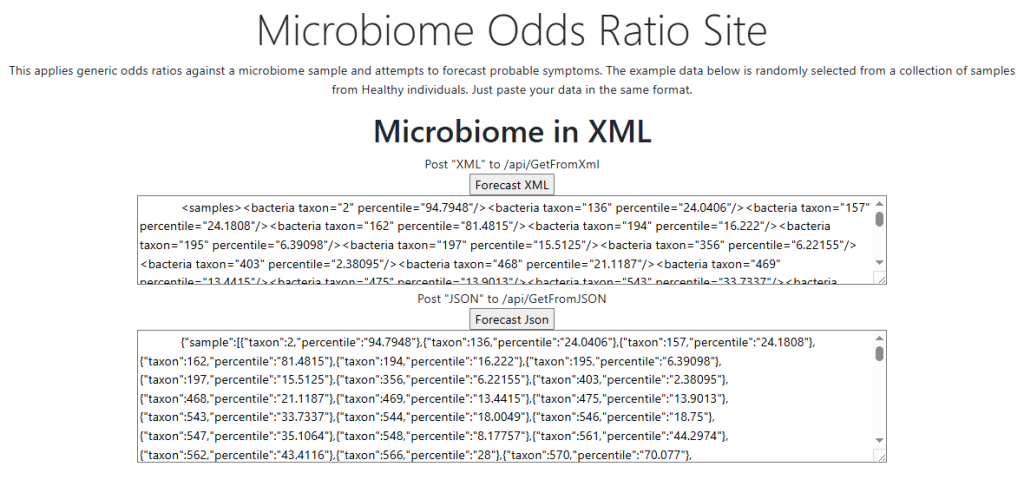
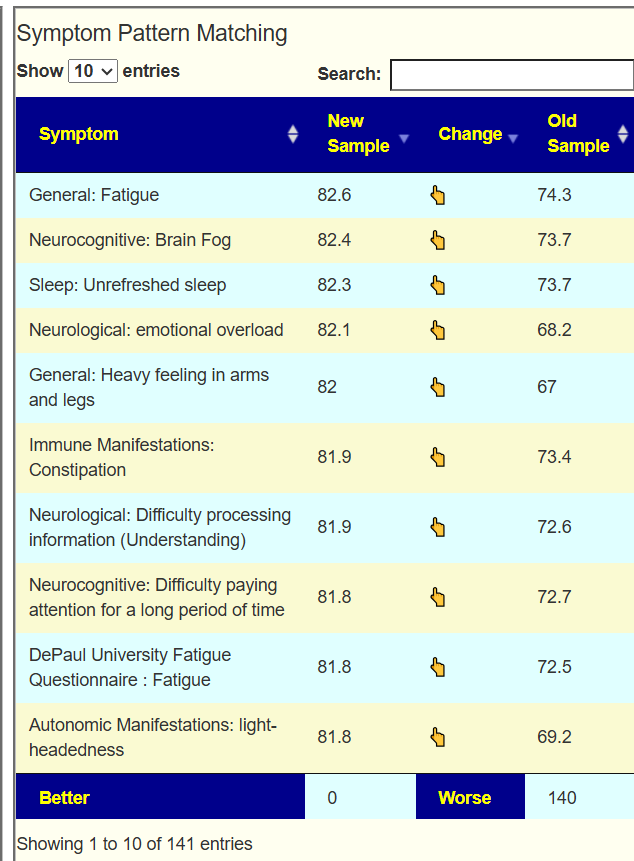
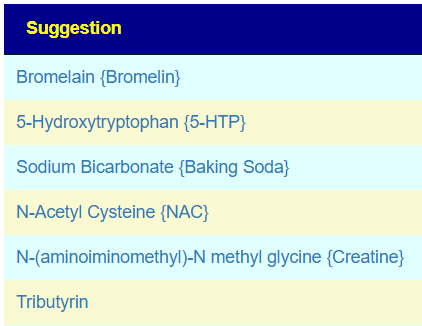
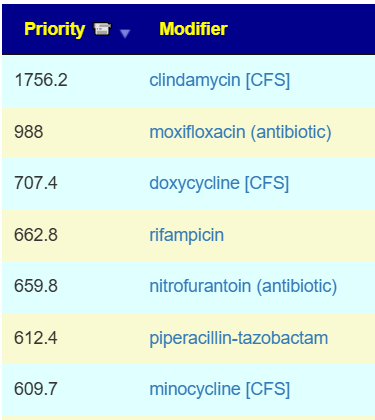
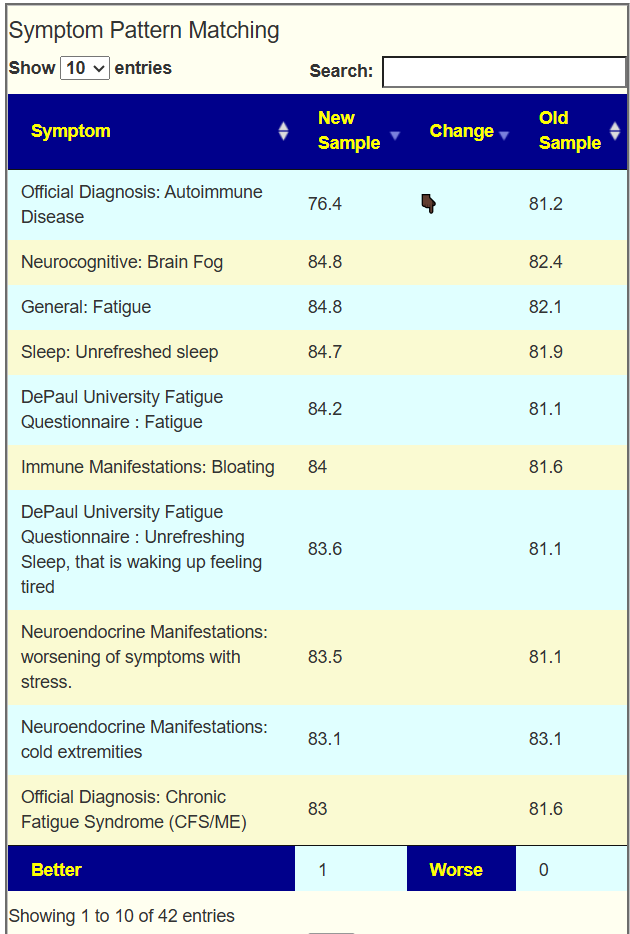
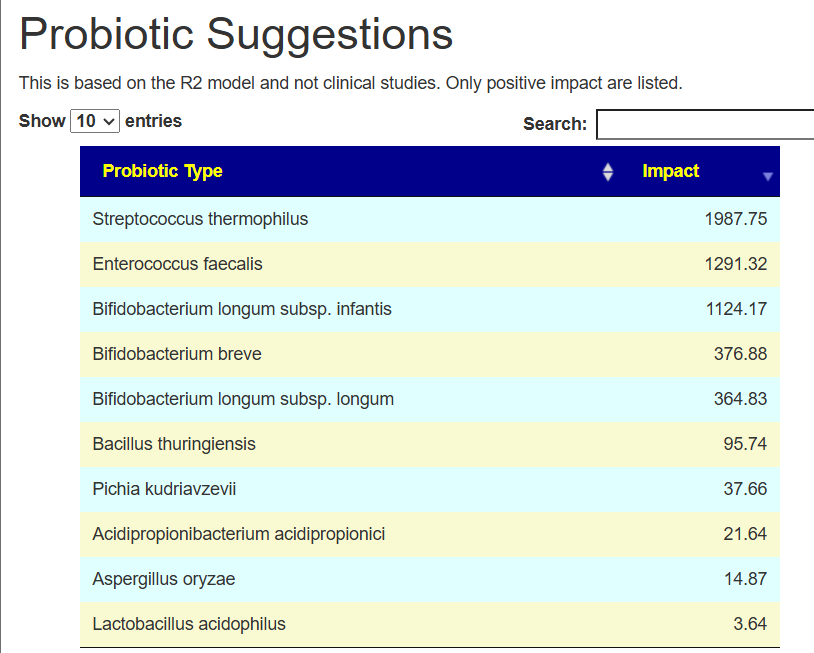
The intent of the Prescription site is to be an adjunct resource for patients. It will generate lists of suggestions based on 14,425,455 facts extracted from 21,279+ studies on PubMed. It may also prepare a detailed list of suggestions with evidence and logic for a MD to review. An example for depression.
Medical professionals do not have time to review an average of 2000 new studies a year. A professional is lucky to review 2 studies a week, not 40. There is a need to alter how they learn.
The site does no use “gathering of hearsay” a.k.a. LLM or ChatGPT, but an older fuzzy logic expert system AI. This is very different from the “hot new AI’s”
I do not know the “right/best way” of determining suggestions. I compute suggestions that mathematics suggests having a greater chance of helping instead of hurting. From feedback and from 2nd sample analysis — it seems to improve people (Analysis Posts on Long COVID and ME/CFS).
The intent is for users to review the suggestions with their medical professionals and get their approval for the plan before starting. Most medical professional will identify any risky items (for example, “Round-Up” once showed up!!) and then say “Whatever, no concerns”
The site is free for individual use. Donations covers operating costs. I have no need to generate revenue from it (50 years in information technology paid well).
Postscript – and Reminder
I am not a licensed medical professional, and the laws where I live prohibit any activity that could be interpreted as practicing medicine or giving personal medical advice. My work is limited to academic and analytical models, and I restrict myself to the language of science and statistics rather than clinical recommendations.
I cannot tell anyone what they should or should not take. Instead, I can present information about items that, based on numerical and statistical analysis, appear to have better odds of improving microbiome-related measures. I am a trained, experienced statistician with appropriate degrees and professional affiliations, and my role is to interpret data—not to treat patients.
All information I provide is for educational and informational purposes only and is not a substitute for professional medical advice, diagnosis, or treatment. Any ideas, rankings, or “suggestions” derived from my analyses must be reviewed and approved by your qualified medical professional before you decide to act on them.
The answers and explanations I provide describe my reasoning and methodology. They are not intended as medical advice for you or for anyone else, and they do not create a doctor–patient or provider–patient relationship. Always consult a knowledgeable licensed healthcare professional before starting, changing, or stopping any treatment, supplement, or health-related regimen.











Recent Comments