I have been asked by a reader to do a review of the latest studies dealing with Salicylate( aka salicylic acid (SA)) Issues. The classic treatment model is to reduce or avoid salicylate foods. Some studies have reported that 2.5% Europeans may suffer from salicylate sensitivity. Aspirin is related and cited as ASA.
My starting point is a 2021 study, Effectiveness of Personalized Low Salicylate Diet in the Management of Salicylates Hypersensitive Patients: Interventional Study which cites some important points”
- How it is cultivated impacts salicylate content.
- The usually recognized as high foods include: legumes (e.g., lentils, beans), vegetables (e.g., cauliflowers, pickled vegetables), fruits (e.g., strawberries, plums, watermelons, raspberries), some cereals (e.g., buckwheat, oat or corn), herbs and spices
- “It was found that the intake of food products with a low glycemic index helps to reduce symptoms in some hyperactive children” [2012]
The personalized low salicylate diet may have a positive effect on reducing self-reported symptoms of asthma, rhinosinusitis, and urticaria, although it is not effective in all patients diagnosed with hypersensitivity to ASA or NSAIDs. The low salicylates diet may be a helpful new tool to support salicylates hypersensitivity therapy, helping to mitigate the symptoms and improve patient well-being. However, further research is needed on the salicylates content of foods, and thus, some modifications of the low salicylate diet may be necessary. Further research is also needed to understand the mechanism of the effect of salicylates in food on the development of food hypersensitivity symptoms.
From Conclusion to Effectiveness of Personalized Low Salicylate Diet in the Management of Salicylates Hypersensitive Patients: Interventional Study
Understanding two forms of salicylates
Looking at amounts in foods, we need to be aware that there are two forms, free (likely to be reacted to) and bound (chemically attached to other things and much less likely to react. Additionally “On analysis, most salicylate-containing foods contain both ASA and SA, and more than one-third contain ASA alone…. ASA challenge reactivity is best regarded as a marker for intolerance to a range of natural salicylates and related dietary phenolics.” [2013]
“
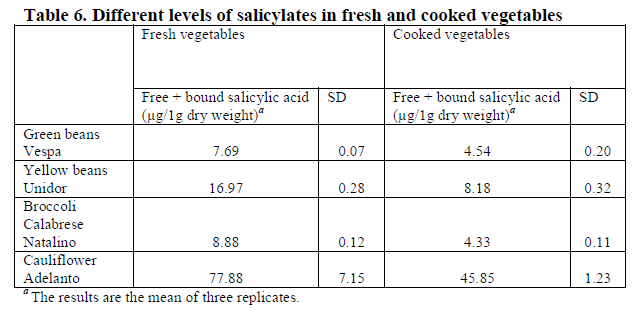
in food available on the European market [2017]
IBS and Mast Cell/Histamine Issues may result
- “These results provide some evidence that food-related salicylates may influence the genesis of symptoms in a subset of patients with IBS. ” [2021]
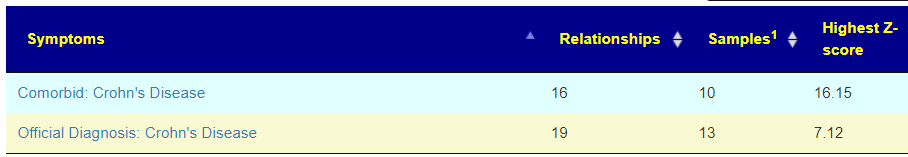
- “Anecdotally, salicylates are said to be a more common trigger compared with other components of the elimination diet…reactions to salicylates can affect different systems and consequently cause different symptoms..2% Crohn’s disease, 7% ulcerative colitis, 6% gastrointestinally mediated allergy…. with rechallenge of salicylates shown to activate mast cells ” [2012]
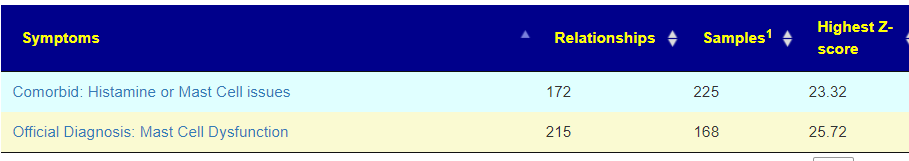
- Idiopathic Mast Cell Activation Syndrome With Associated Salicylate Intolerance [2018]
- Aspirin activation of eosinophils and mast cells: implications in the pathogenesis of aspirin-exacerbated respiratory disease [2014]
- “double-blind placebo-controlled challenges identified problem foods in 6% to 58% of [IBS] cases. Milk, wheat, and eggs were most frequently identified to cause symptom exacerbation; of the foods identified the most common trait was a high salicylate content.” [1998]
- This was an interesting study dealing with Mast Cell activation : “Postmortem diagnostics of assumed suicidal food anaphylaxis in prison: a unique case of anaphylactic death due to peach ingestion [2021]”
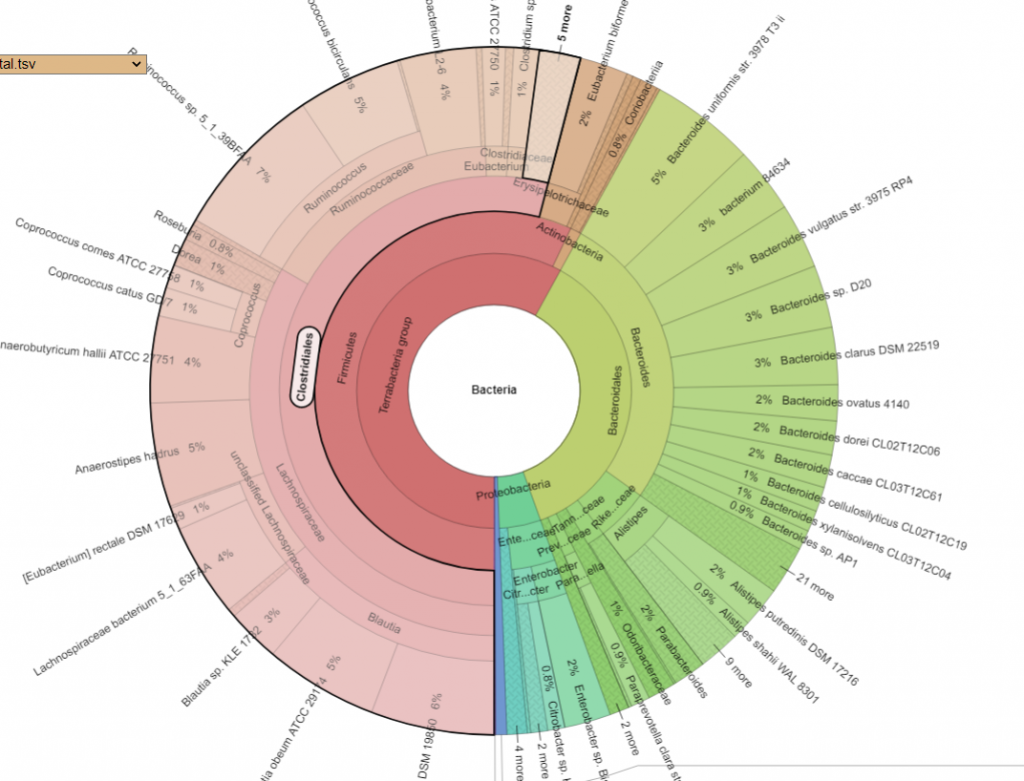
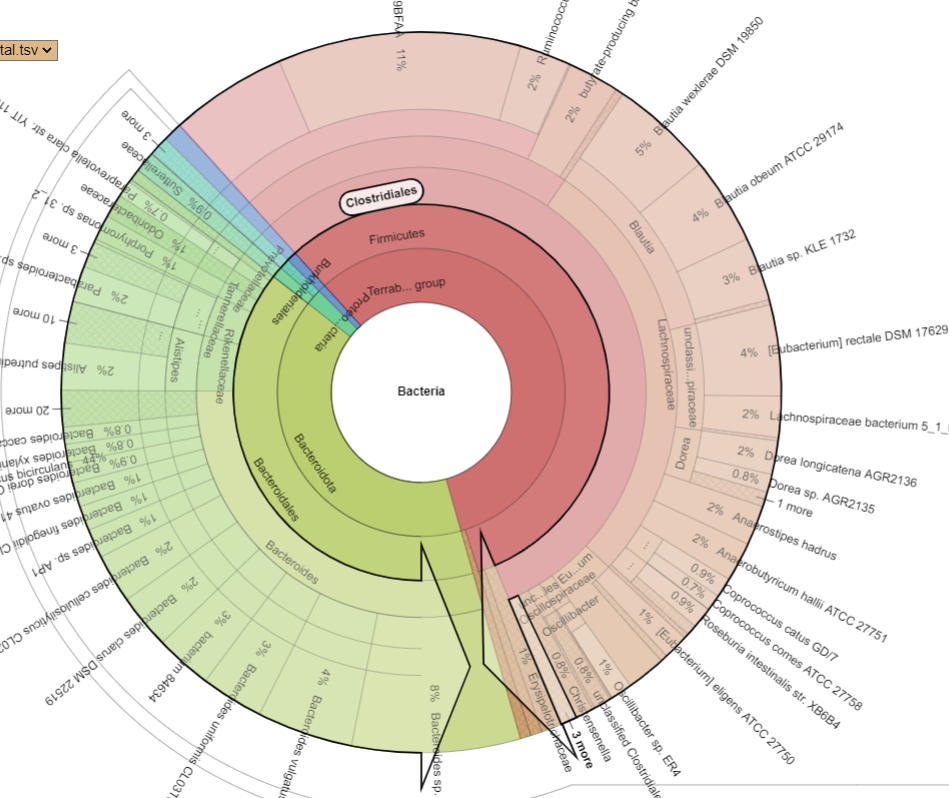
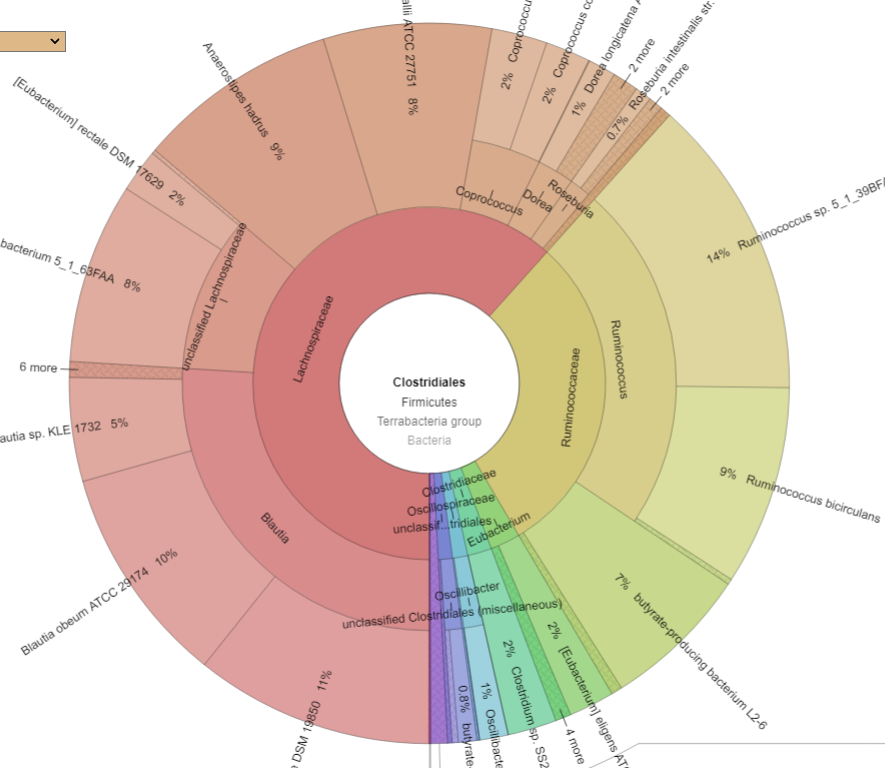
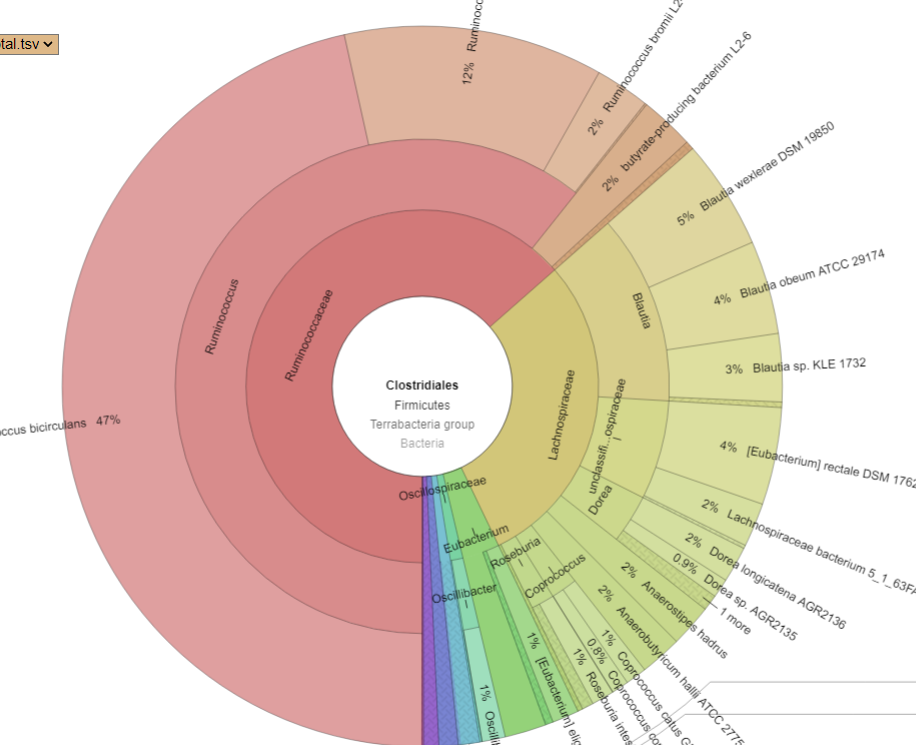
Bacteria Associated with Salicylates Sensitivity
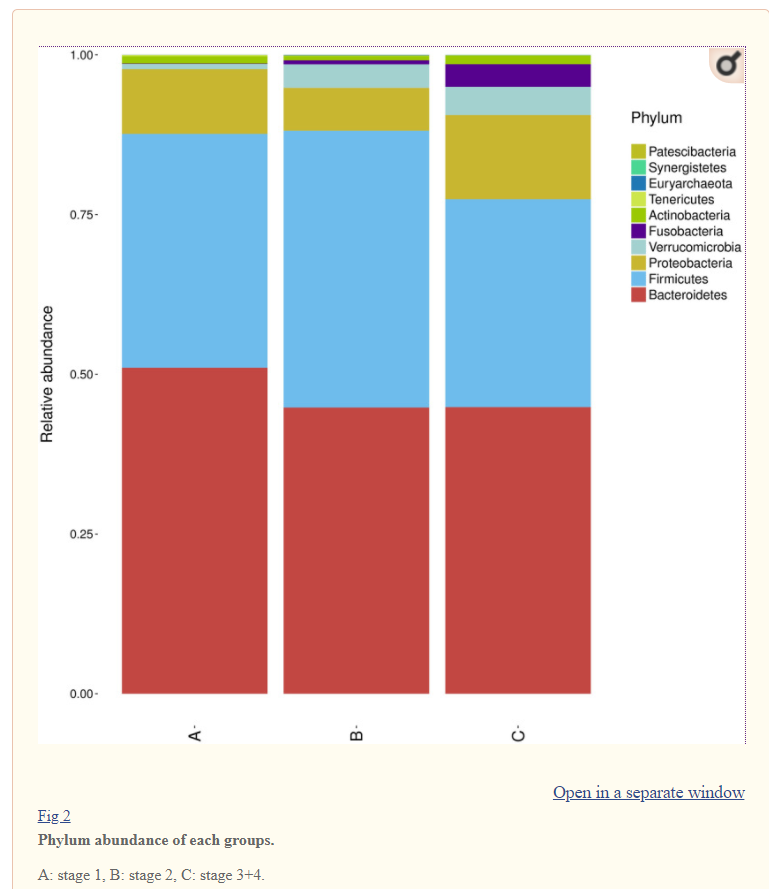
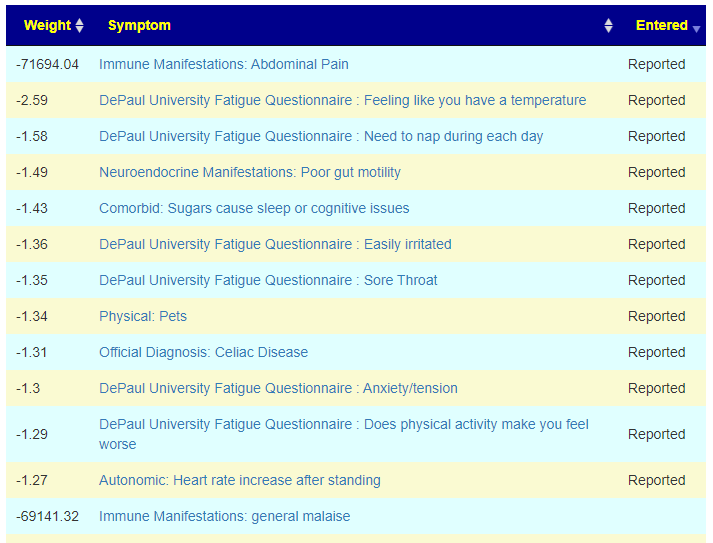
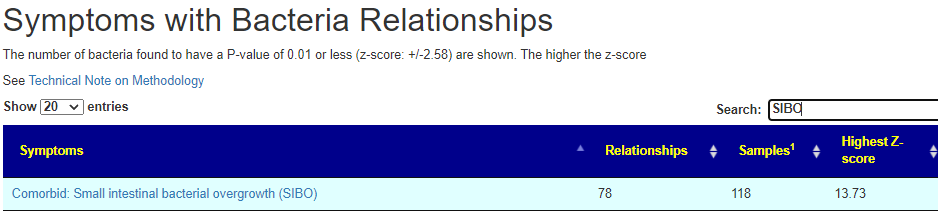
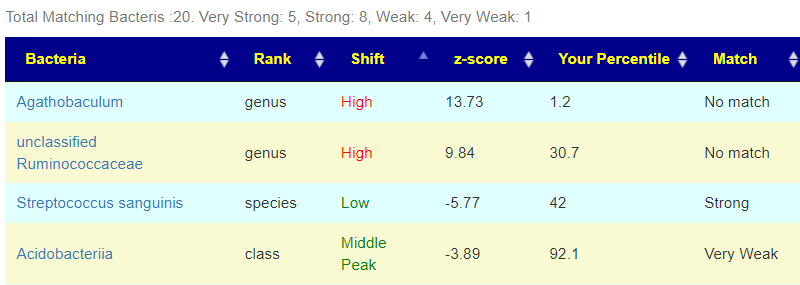
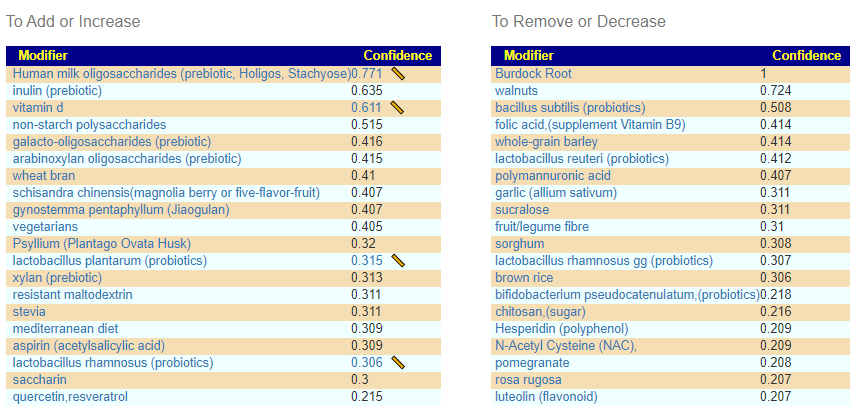
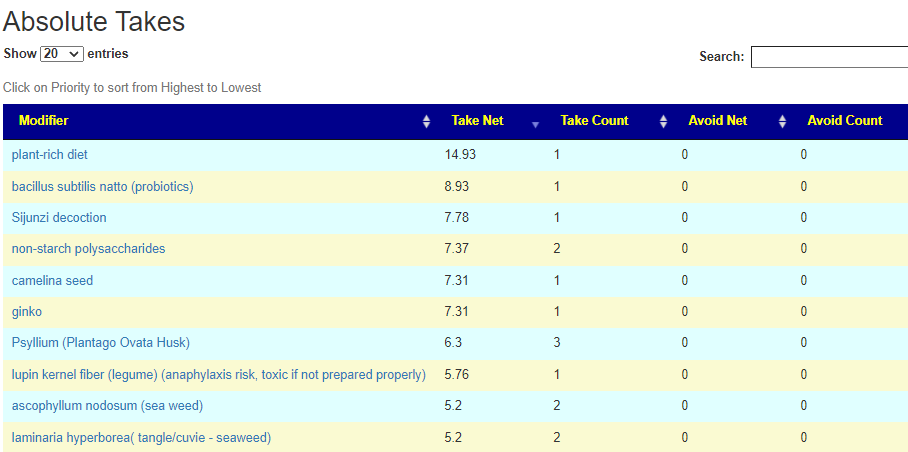
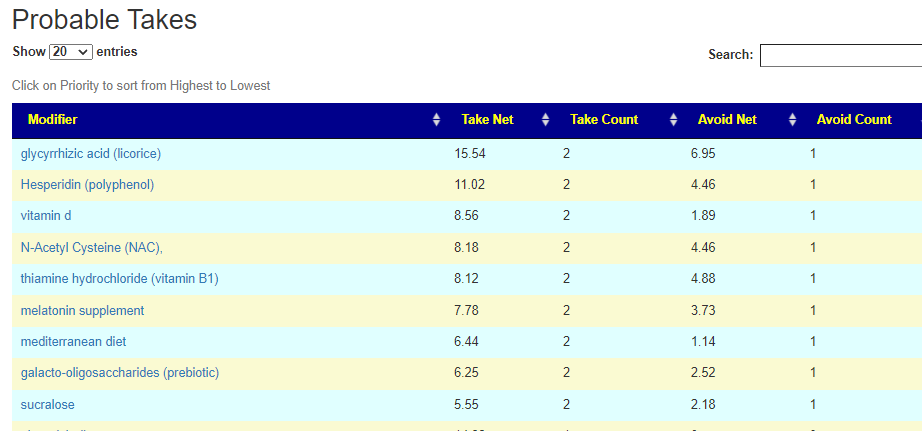
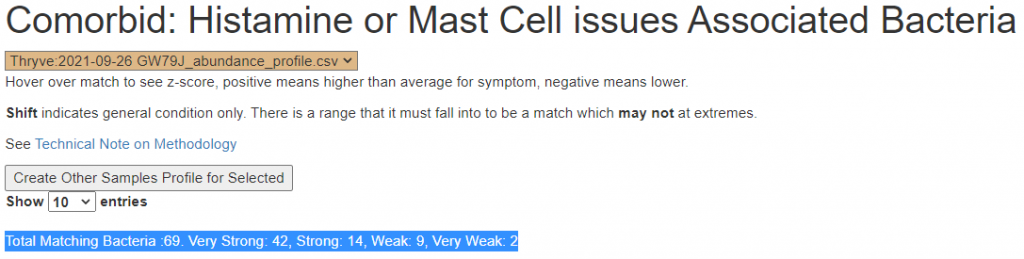
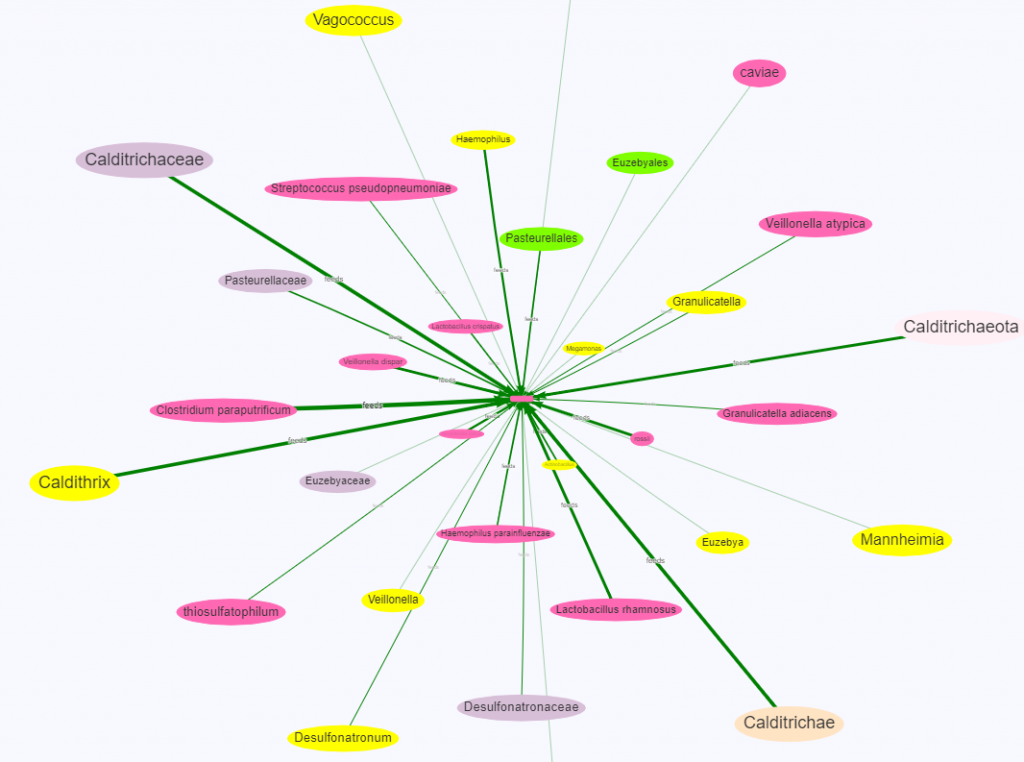
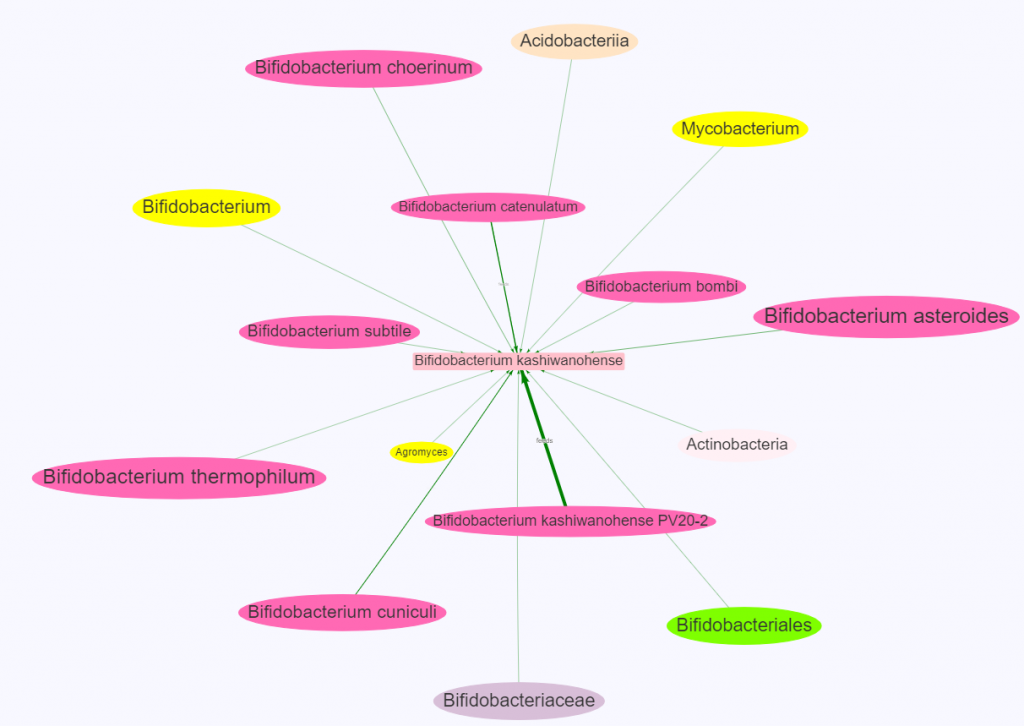
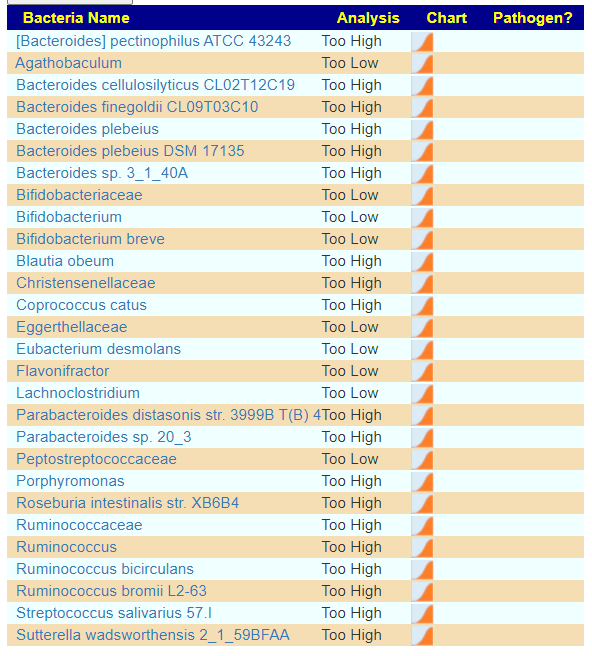
There is nothing on PubMed – no one has done a study of the bacteria shifts seen with this condition. Fortunately, Citizen Science on Microbiome Prescription found some shifts: Comorbid: Salicylate sensitive The number of samples is low, so hopefully this will improve. The most likely bacteria are:
- LOW
- Bifidobacterium subtile (unfortunately, not available as a probiotic)
- Actinobacteria
- HIGH
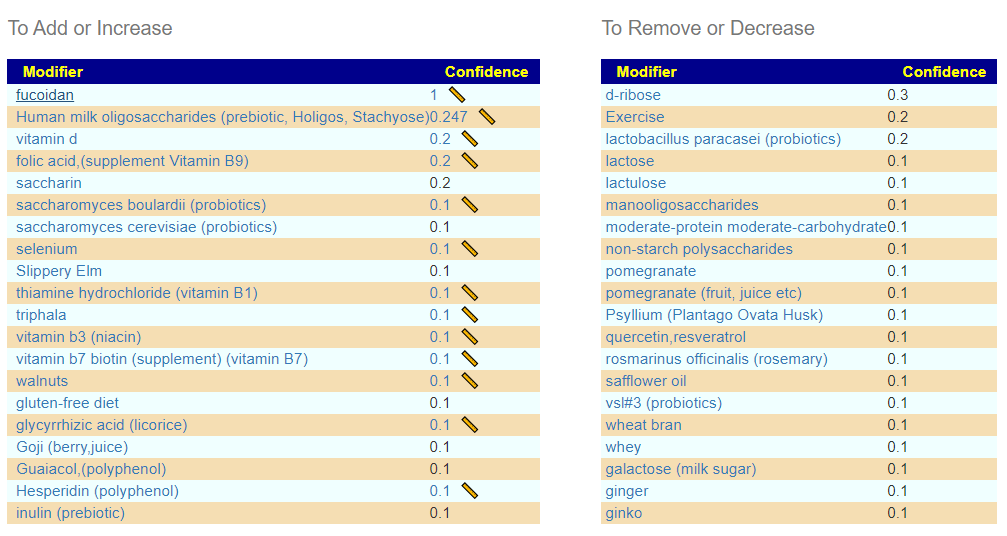
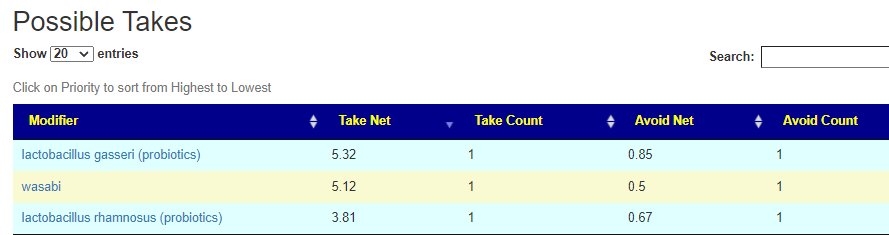
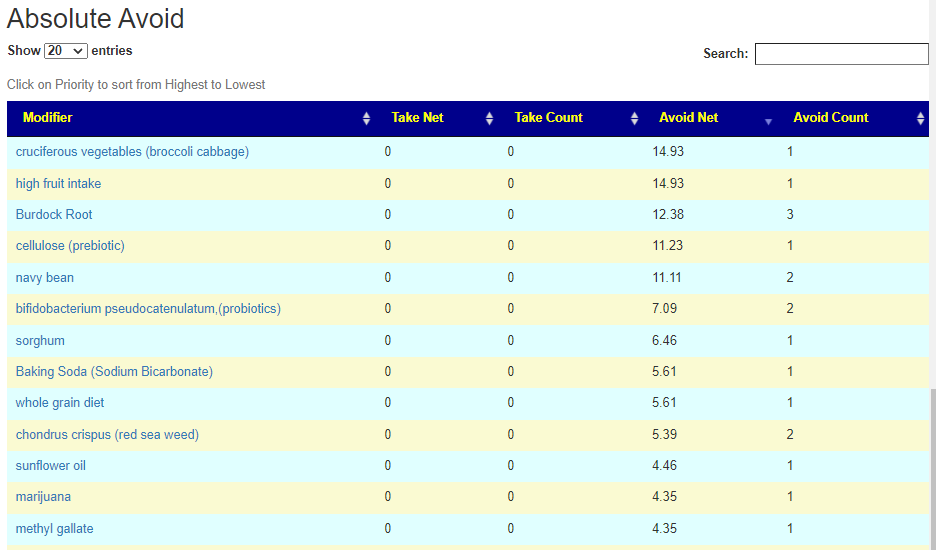
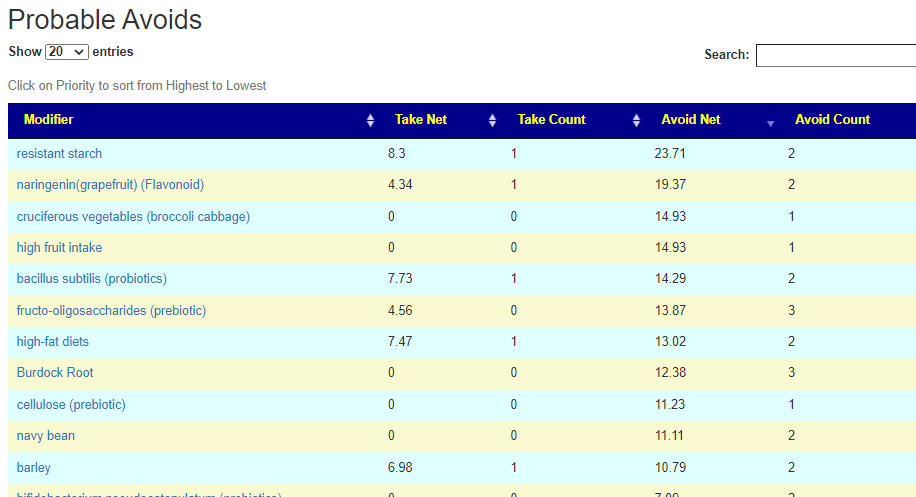
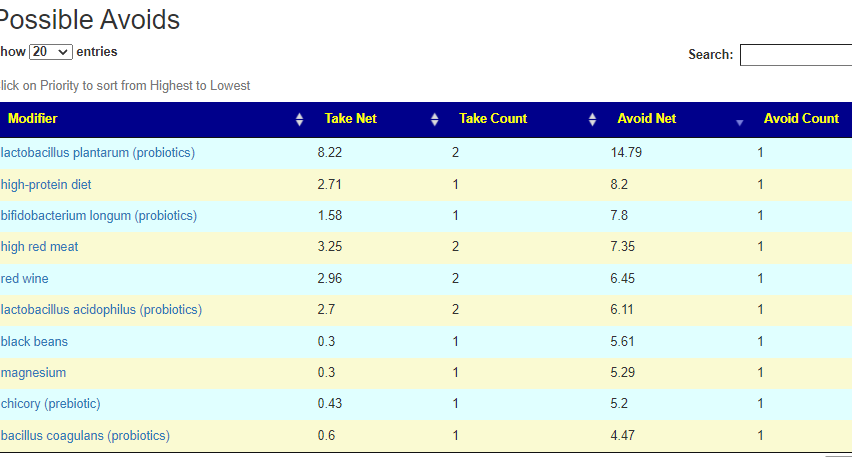
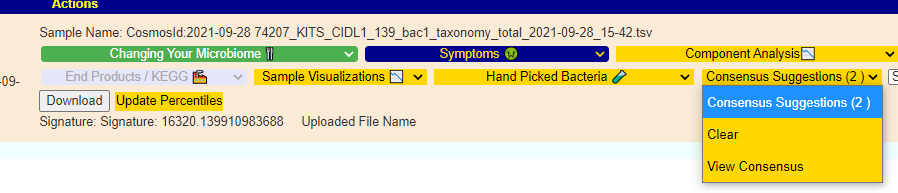
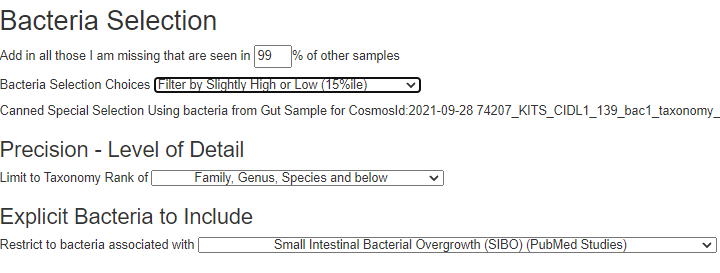
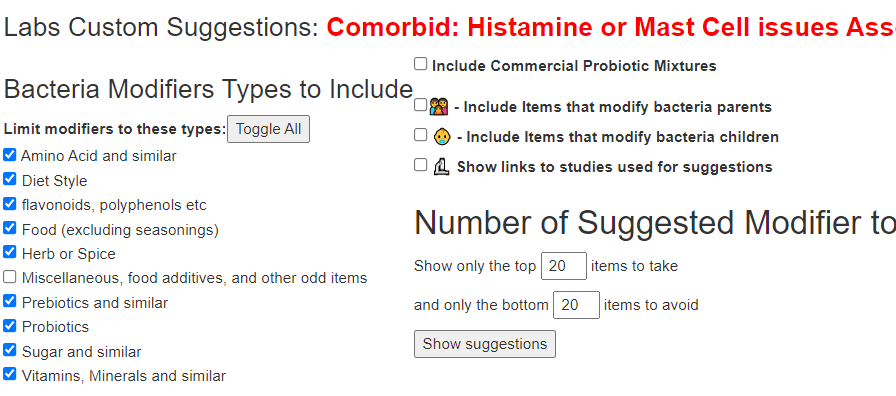
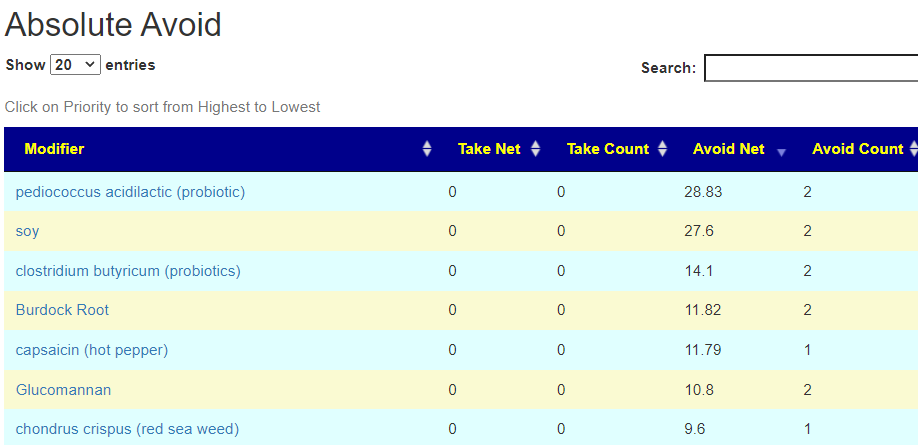
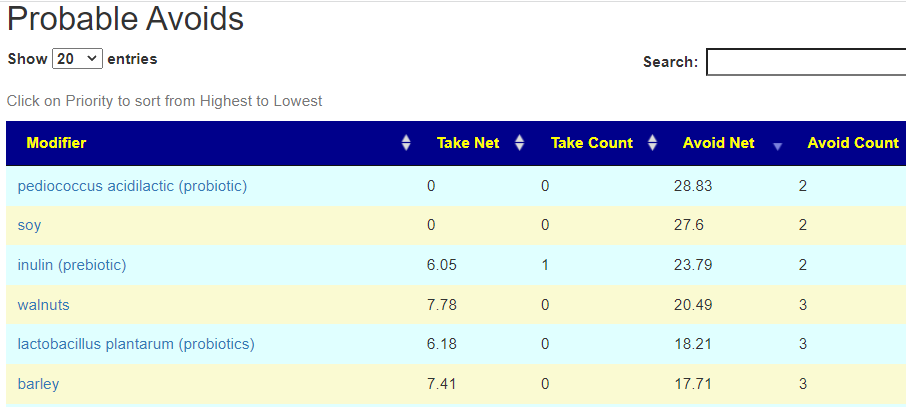
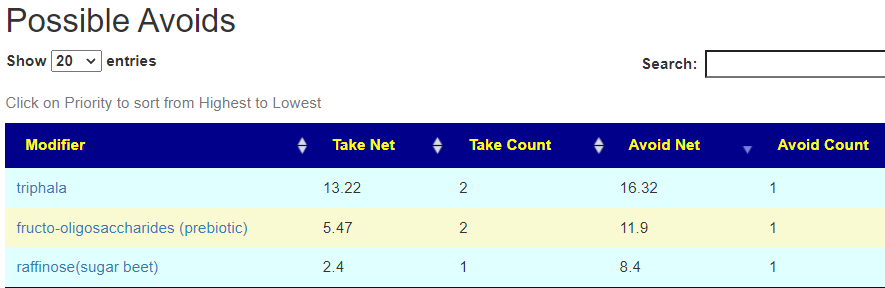
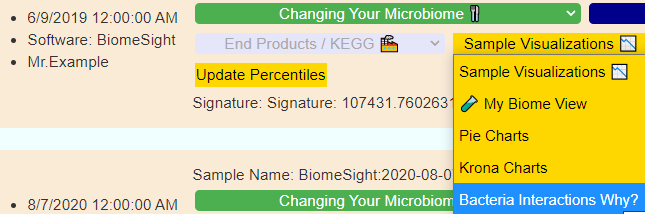
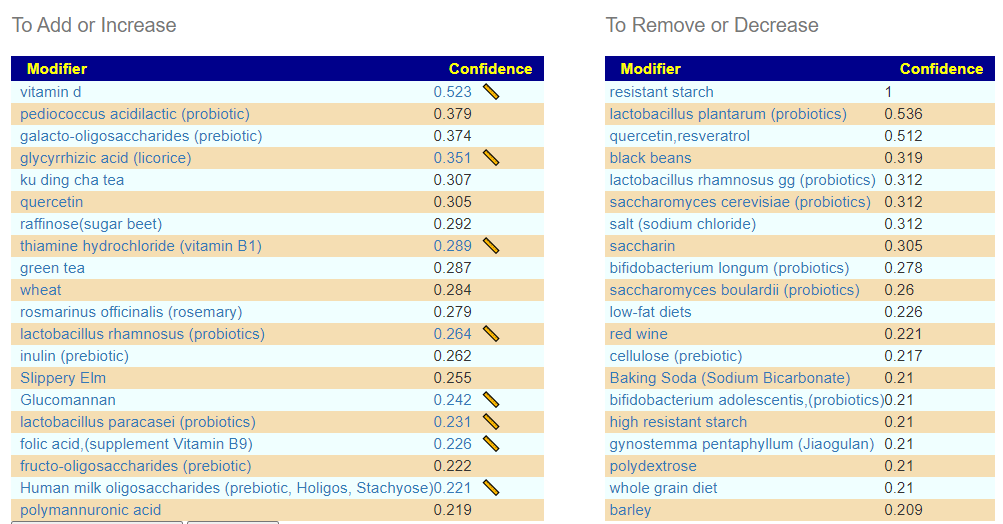
If you have a salicylate sensitivity and a 16s lab, you may wish to upload it and see what is suggested. Here is an example walk thru
Salicylates Lookup Table
From a variety of studies on PubMed, I extracted various measurements — always going with the highest value (for those that are sensitive, the safest).
- Which Fruits and Vegetables Should Be Excluded from a Low-Salicylate Diet? An Analysis of Salicylic Acid in Foodstuffs in Taiwan [2018]
- Salicylates in foods [1985]
- The overall content of salicylic acid and salicylates in food available on the European market [2017]
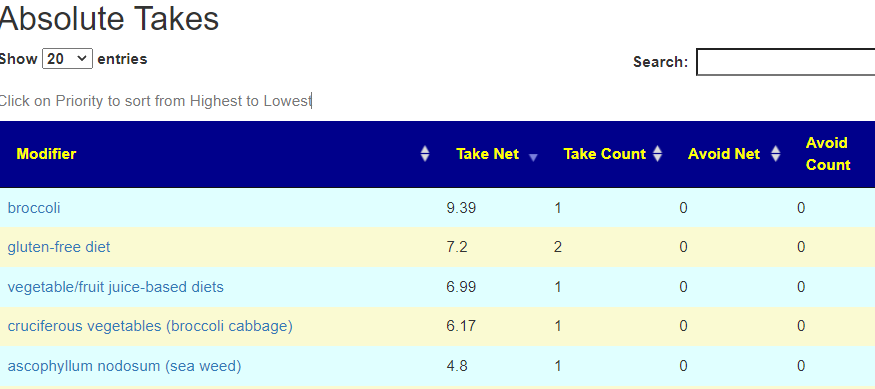
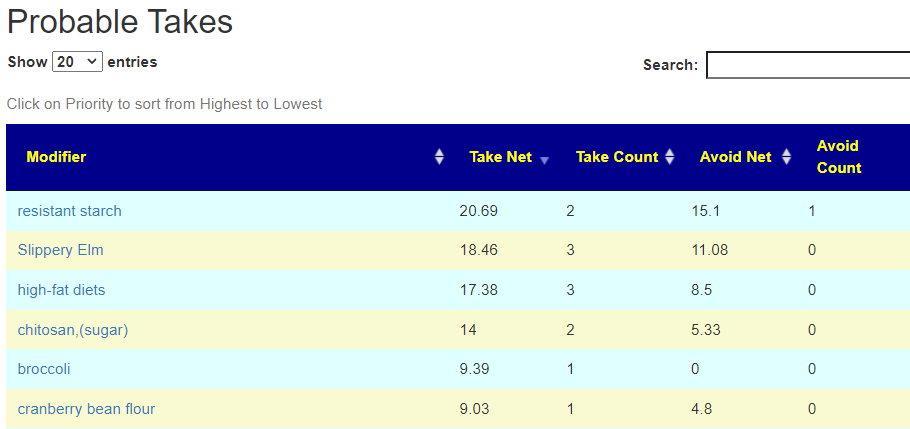
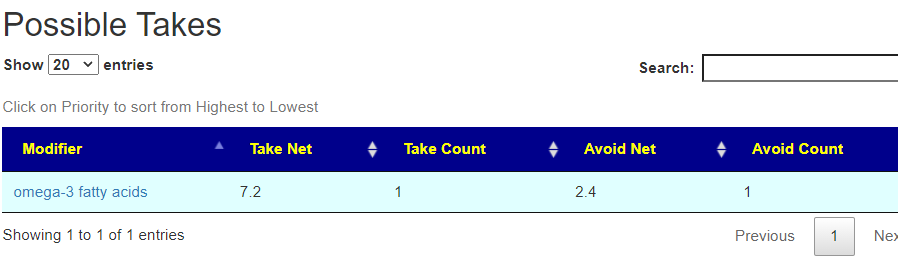
On line table.















Recent Comments