One of the common misconception is that there is a “normal” microbiome that can be used as a reference. Below is a chart from “Metagenomic sequencing of fecal DNA“. Diet makes a major impact on the distribution and volume of the bacteria. Firms like uBiome show the pattern of people with different diet on their site as shown below:
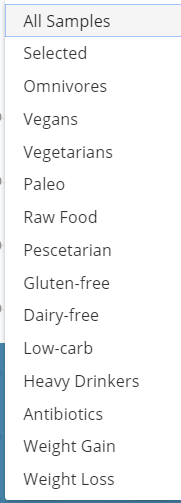
This means that interpreting the microbiome is subjective and complex.
- “In a study of gut bacteria of children in Burkina Faso (in Africa), Prevotella made up 53% of the gut bacteria, but were absent in age-matched European children.”[2010]
The chart below is for healthy individuals in 12 different countries. In some cases neighboring very similar countries (Sweden [SE] and Denmark [DK]) have very different compositions.

This great variation means that testing the microbiome can only be done as group of individuals living in the same area with similar eating habits and state of health.
For more information, see this 2017 post on CFSRemission.
Art of the Microbiome and not the Science of the Microbiome
The above information means that any and every attempt to have definitive scientific rigor with your microbiome will fail to pass the highest standards. How can we possibly tell that the issue is connected with a disease and not diet, or even DNA!!!
I have build Fuzzy Logic Artificial Intelligence system since the late 1980’s. We may not know what the definitive answer is, but we can conclude more probable answers than ignoring what we read in studies.
Almost every study comes down to association. There may not be any actual direct links or causation.
For instance, cheese consumption in the US between 2000 and 2009 correlated with the number of deaths by entanglement of people in their bedsheets [2]. The number of people who drowned in a pool between 1999 and 2009, correlated with the number of films starring Nicholas Cage that were released during that time [3]. Does one of these factors cause the other? Probably not.
https://www.students4bestevidence.net/association-is-not-the-same-as-causation/
In general, we will assume from the studies that there is some connection – for example, taking clostridium butyricum (Miyairi) probiotics has been reported to increase lactobacillus acidophilus levels. The actual mechanism may be complex: Miyairi may inhibit another bacteria that inhibits the lactobacillus acidophilus. If we put Miyairi in a yogurt that only contains lactobacillus acidophilus, we may find that it decreases the level … because the Miyairi steals food from the lactobacillus acidophilus.
This type of complex situation is what fuzzy logic attempts to control.
Under this page you will find pages:
- Describing my preferred method to identify problems with a random person microbiome.
- Describing what is meant by Condition Profiles on https://microbiomeprescription.com/
Choices for Determining What you wish to change
Once you have logged on, if you select Advanced Suggestions
https://microbiomeprescription.com/email/samples
You will go to the custom suggestions page. On this page, you will see your samples and links for Bacteria Selection choices.
http://microbiomeprescription.azurewebsites.net/data/CustomSuggestions?sampleid= ….
There are multiple paths available for determining what could be wrong, and thus what you may wish to modify. Click on the above high lighted link will show you the actual bacteria picked from each of these methods. A short summary of each is below:
- Bacteria that has an association with symptoms that you have. This will be the smallest list — but it is the most certain of being right
- Bacteria whose population are outliers (that is far away from the expected patterns). This is the medium size list — it is uncertain if they are significant
- Bacteria above or below normal ranges. This is the simplest to understand and may be most leading. The typical 16S report has 500 to 1500 bacteria listed. Assuming the normal range is the middle 90% (5% high and 5% low), we would get 50 to 150 bacteria listed here by random chance.
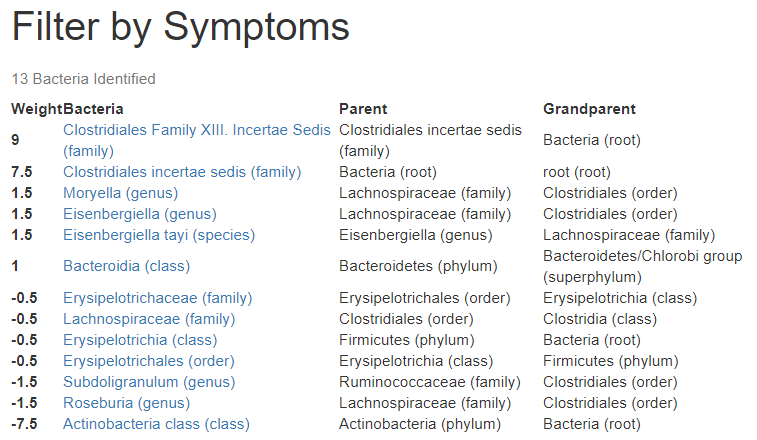
I did some counts on different samples, which confirms the counts from each method.
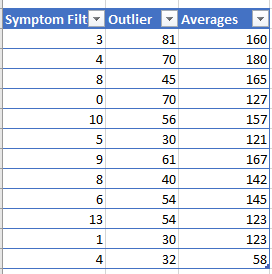
Note: The symptom and outlier filter is dynamic. As more data is added, their ability to detect improves. For symptoms, there must be 16 people people reporting the same symptom before we will do detection of association.
In Summary
You can try all three methods, and experiment. You really want to get the smallest list of bacteria (because in figuring out suggestions, the more bacteria you include, the more complex it gets!).
For myself, I do suggestions from each and found a few items appeared in common with all of them.
Recent Comments