I am newly ill from ME/CFS (4-5 months) and wonder if my results look like a “typical ME/CFS” for you, or Long Covid for that matter. I am a bit confused and some of the recommendations are contradictory.
From Reader. He used Biomesight.com (serves the world, discount code “Micro”)
This will be an interesting analysis — the microbiome evolves over time, so a 20 year ME/CFS and a few months of ME/CFS will have differences. I know of no clinical studies looking at “fresh ME/CFS”. There are studies for “fresh Long COVID”.
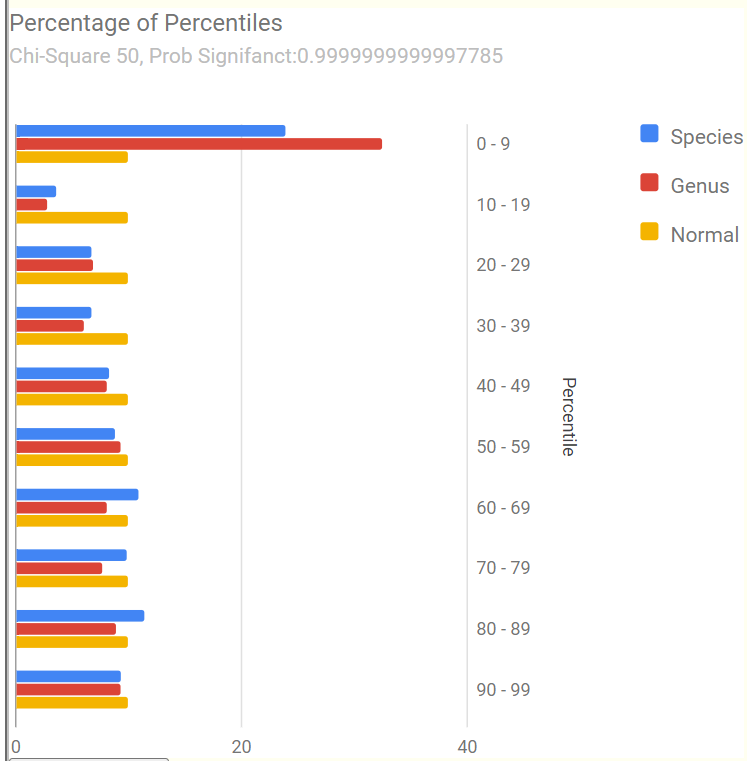
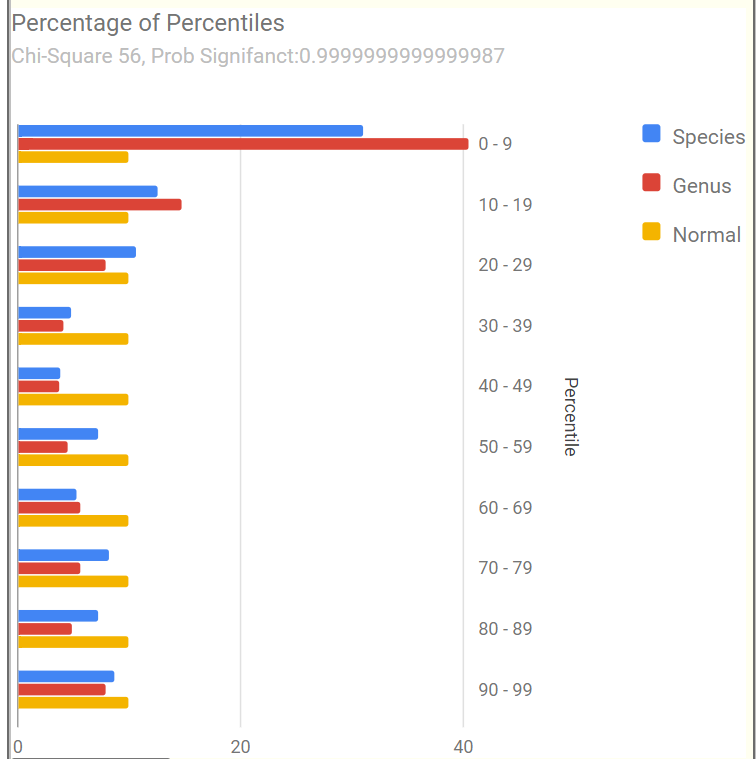
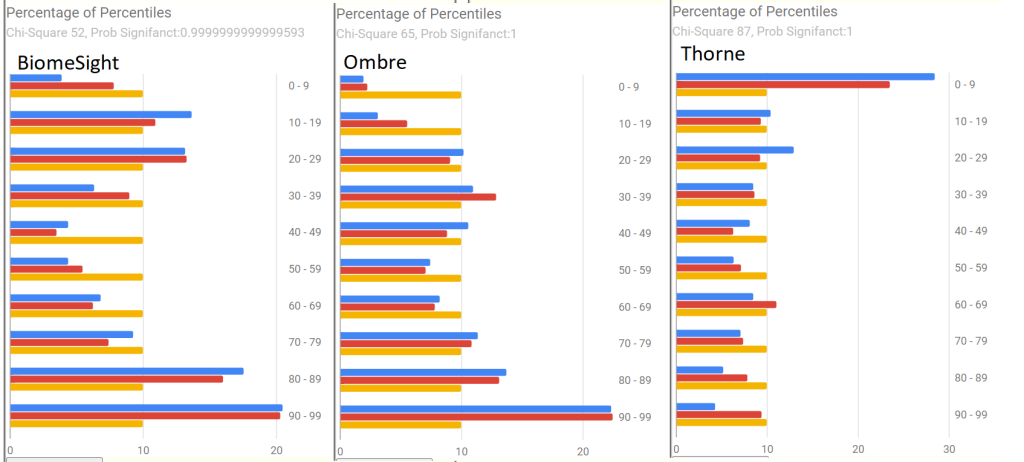
Percentages of Percentiles
This tend to have a regular pattern for ME/CFS and Long COVID for most people. Over representation of bacteria in the 0-9% percentile range. This matches his pattern. A healthy person would have all of the bars around 10% – they are not.
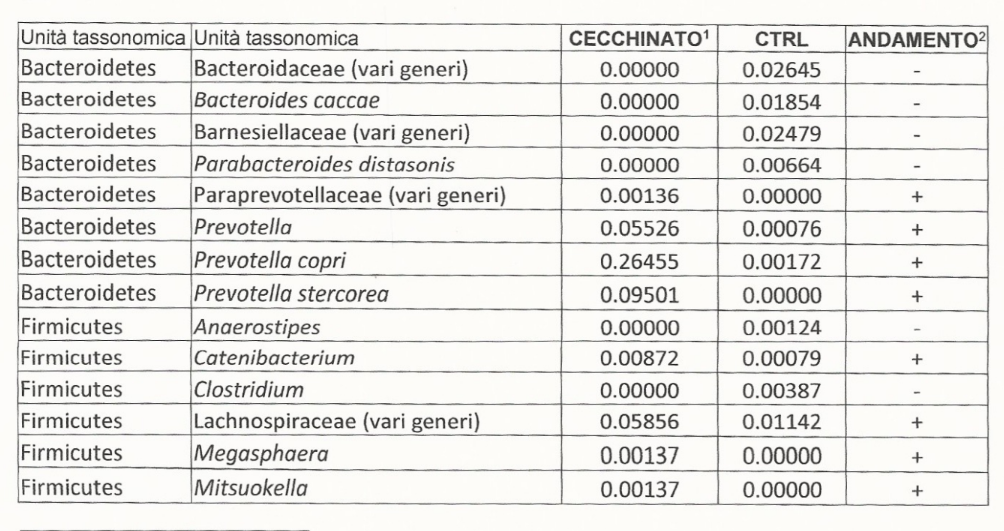
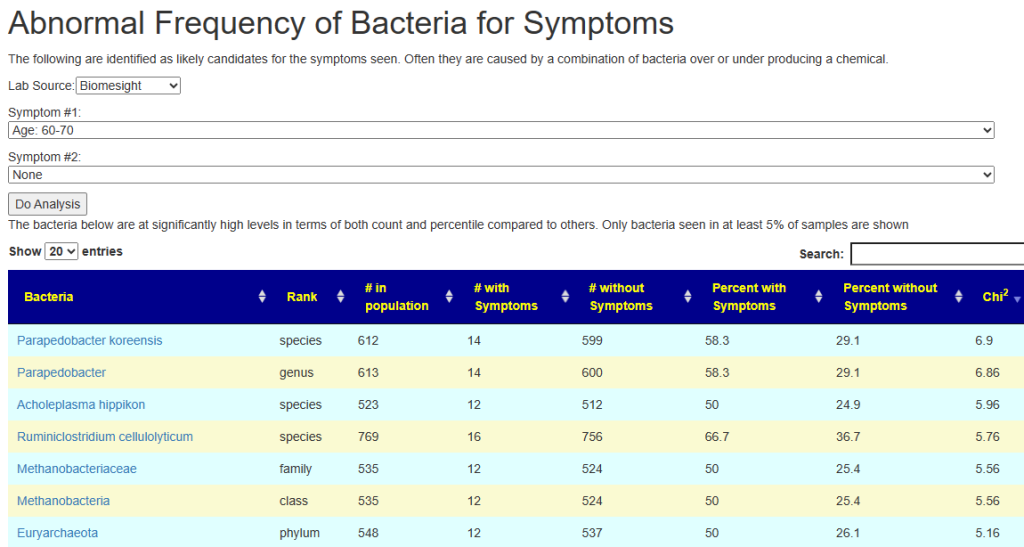
Going to the Potential Medical Conditions Detected, there was nothing significant. Prevotella copri is 2% (78%ile) which is borderline for mold issues — it would be good to do inspect the home for that risk. Dr. Jason Hawrelak Recommendations comes in at 99.7%ile with high Methanobrevibacter and low Bifidobacterium being the most severe shifts. Anti inflammatory Bacteria Score is 63.1%ile which is a bit better than most people. So, many medical professionals would tell the patient that I cannot see anything wrong.
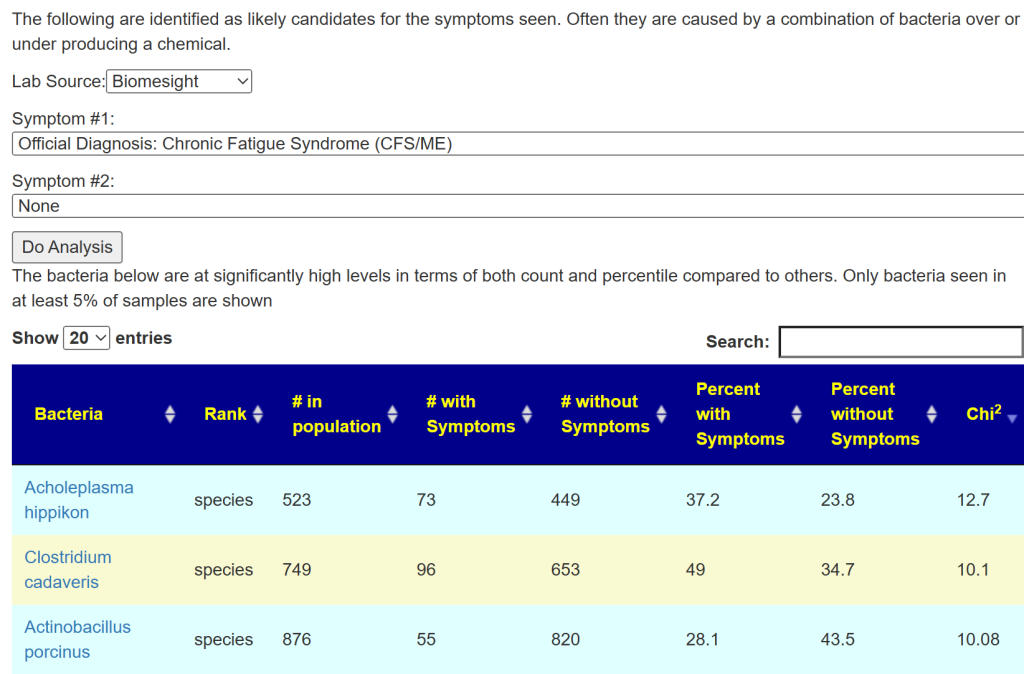
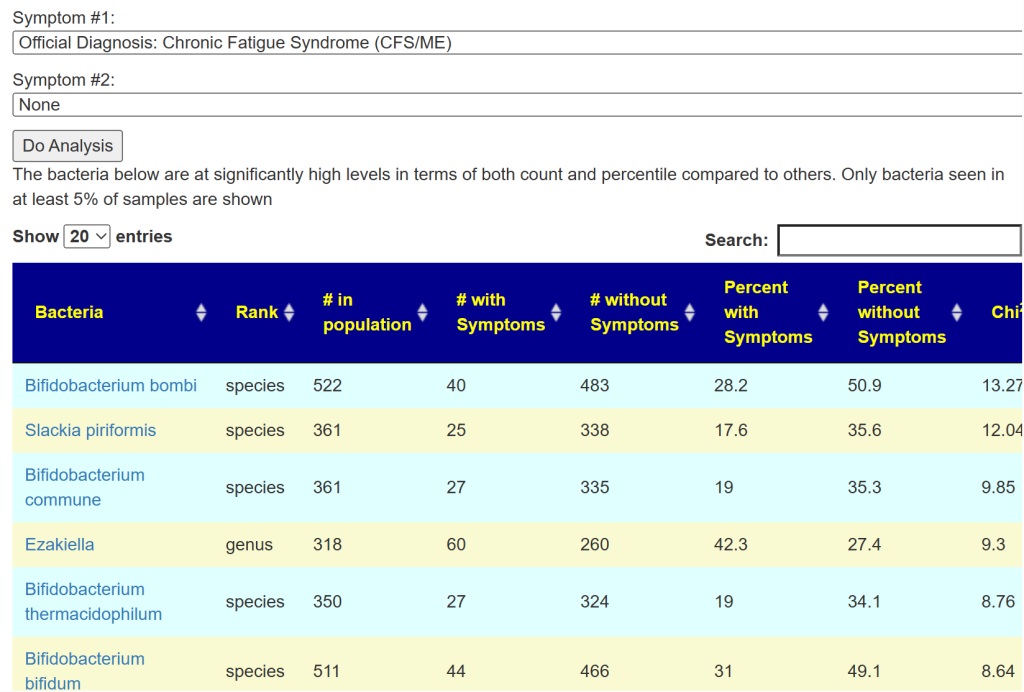
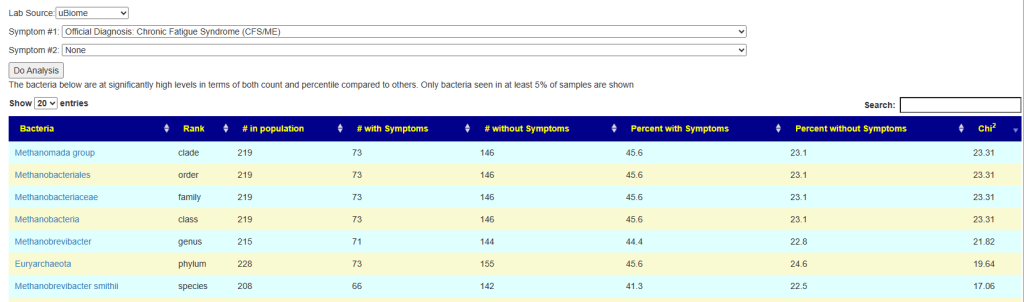
Special Studies
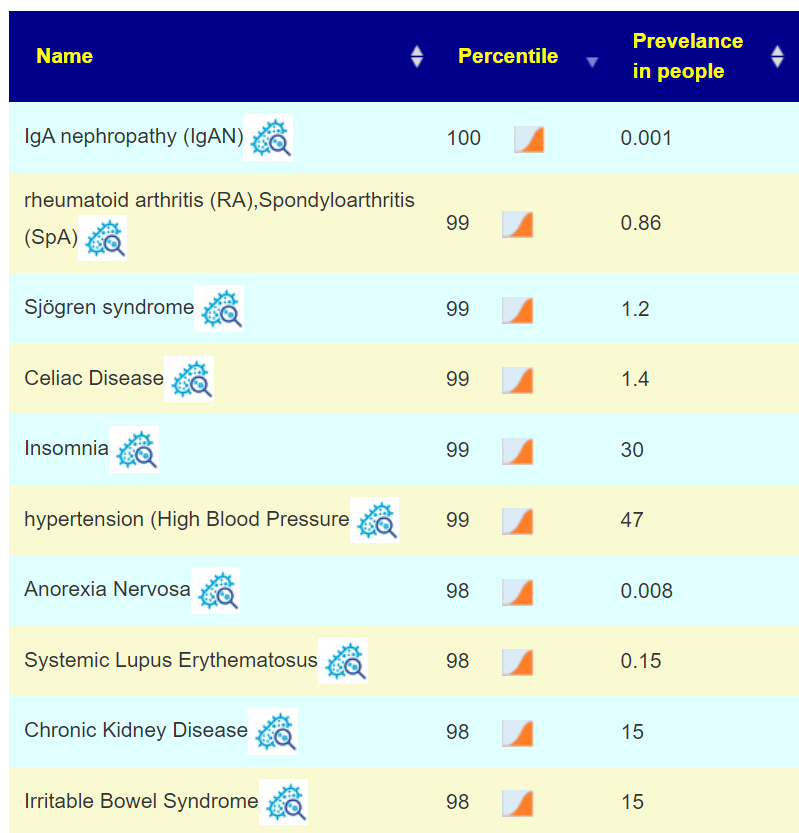
Special studies are statistical studies that uses samples uploaded and look at the self-reported symptoms. The analysis is done individually for each lab (needed because of differences in how labs process samples). In this case… we see that COVID Long Hauler is by far the strongest match, almost double the next one.

I did the usual Just Give Me Suggestions and then did the Long Hauler suggestions . This gave five (5) packages of suggestions. I then looked at the antibiotics suggestions, focused on those used with ME/CFS. The top ones are:
- METRONIDAZOLE (ANTIBIOTIC)
- AZITHROMYCIN,(ANTIBIOTIC)S
- CHLOROQUINE DIPHOSPHATE
- DAPSONE (ANTIBIOTIC)
If you have a cooperative MD, I would suggest following the protocol of the Belgium MD, Cecile Jadin: Dr. Jadin’s Current Protocol for ME/CFS.
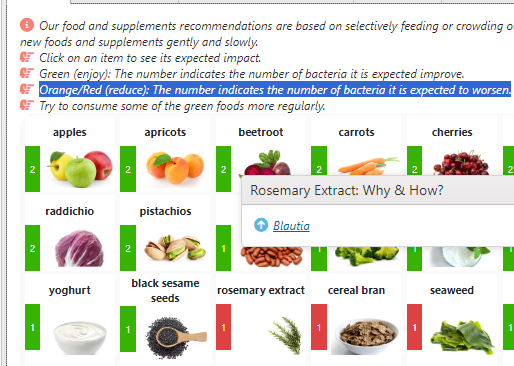
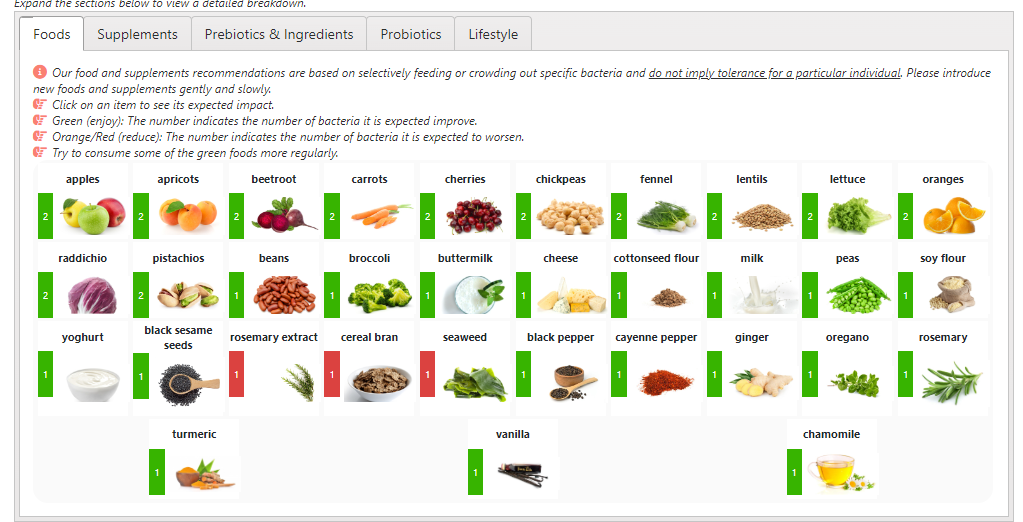
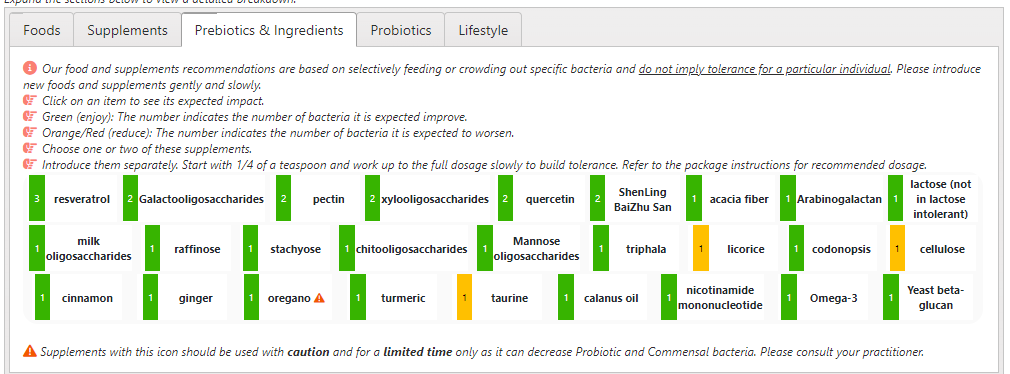
The list of suggestions to take is actually bigger than usual.
- REMEMBER: There is no need to take all of them, just take what works for you (i.e. no adverse effect and acceptable cost).
- Probiotics should be rotated (change to a different one) every 7 to 14 days. Probiotics often work by producing natural antibiotics. Continuous taking of the same probiotics may result in it not working because of “natural antibiotic resistance”.
- The colors have no meaning except as indicators for category. For example, green is probiotics
- Dosages are those that have been used in clinical studies (for other conditions), and thus deemed safe dosages. Often I have see people taking < 1% of these dosages and wondering why nothing happens.
For example, it you are lactose intolerance, then ignore the lactose suggestion. If not, regular cups of good Belgium Cacao would be a good prescription!

The safe retail probiotics were calculated to be
- symbiopharm/ symbioflo2 – an E.Coli probiotic from Germany
- Filmjölk (SE) / Filmjölk – a Swedish milk drink
- enterogermina – Bacillus Clausii
- SunWavePharma / Spor Sun – Bacillus Clausii
Why are these lists not the same? The latter list are ones that will not shift a single one of the bacteria we are focused on in the wrong direction: NOT A SINGLE BACTERIA. This is an extremely safe conservative suggestion. The top list with probiotics in green often contain probiotics that shifts 30 bacteria in the right direction and 3 in the wrong direction. The odds are that they are very likely to help.
We have a third list of probiotics (to make probiotic suggestions even more confusing), the KEGG suggestions. This looks at what enzymes your microbiome are low in, then sees which probiotics can provide those enzymes. Our goal is to reduce enzymes starvation; this cascades in metabolites — chemicals that the body uses — starvation. The key items from this list are:
- Escherichia coli (symbioflor-2 above or Mutaflor)
- Bacillus subtilis subsp. natto (1/3 of contribution of E.Coli probiotics)
- Bacillus subtilis (1/3 of contribution of E.Coli probiotics)
- Lactobacillus plantarum subsp. plantarum (1/4 of contribution of E.Coli probiotics)
If you have significant brain fog, I would be careful with taking lactobacillus probiotics. Some retail species can increase brain fog.
The Avoids
Frequently “good suggestions for general health” are bad for some conditions. The avoid list of things to avoid is short

Some quick translation: no iron supplements (ferric citrate), beta-glucans usually means no oats, barley or Reishi mushrooms. The names are those used in clinical studies — so they tend to be “unresolved” often in common speech.
Suggested Cycles
After implementing the above suggestions for 6 weeks, do another test and see what has changed.
Questions and Answers
Q: Is it best to take the antibiotics + the protocol you suggested ? Or is it one or the other?
- A: My own choice would be to do both at the same time. If you follow Jadin’s approach, then
- First week of antibiotics per month — no probiotics at the same time
- Second week do some of the probiotics (I am inclined towards the E.Coli probiotics but that is based on my personal experience — your mileage may vary)
- Rotate the other supplements over the weeks. I would suggest 10 days and then change to a different set.
Q: And lastly, in the « Avoids », the « vegetarians » suggestion is a little confusing to me. What does it mean?
- Vegetarians mean no animal or fish is being consumed. So, have fish — but some animal proteins are to be reduced: no pork, moderate beef. Duck, chicken and rabbit are fine.
Q: The suggested dosage in the suggestions seems very high. 30g of Vitamin C per day??? Is this correct ?
- The actual dosage should be discussed with your medical professional. The dosages are the highest that have been done in clinical studies and thus assumed to be safe dosages. We have no data on what the threshold for an effective dosage should be. I have seen a few studies where 1000 mg of a substance has minor/no effect while 1100 mg has twice the impact and 1200 mg has four times the effect. For many substances there seem to be a threshold that triggers changes.
- Usually start at 1/8 of the dosage and double it every second day until there is a response. If very good keep at that dosage. If bad, cut the dosage in half and try a few more days. Give the body time to adapt.
My comment about mold caused him to check his environment carefully…. what he found out of sight!
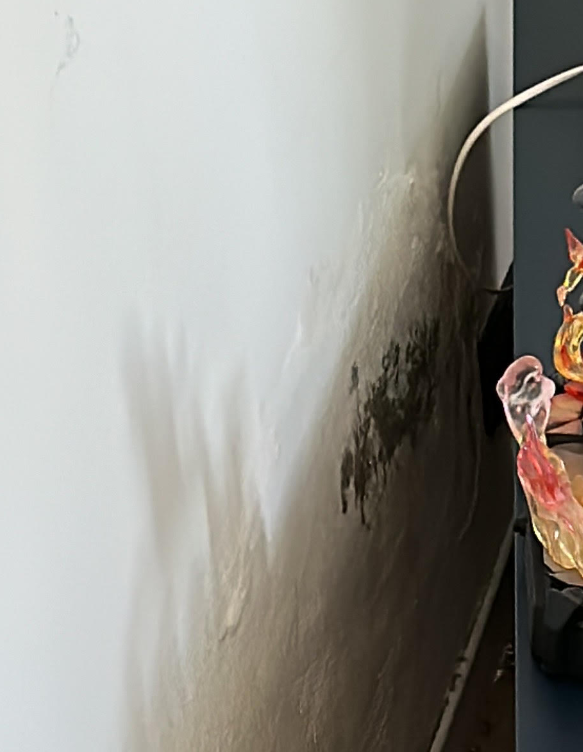

Mold and ME/CFS Relationship
“Urine specimens from 104 of 112 patients (93%) were positive for at least one mycotoxin” from Detection of Mycotoxins in Patients with Chronic Fatigue Syndrome [2013]
“Environmental factors – exposure to mold or toxins has been suspected as a trigger for ME/CFS. However, associations of specific environmental factors with ME/CFS have not been established.” [CDC]
Postscript – and Reminder
I am not a licensed medical professional and there are strict laws where I live about “appearing to practice medicine”. I am safe when it is “academic models” and I keep to the language of science, especially statistics. I am not safe when the explanations have possible overtones of advising a patient instead of presenting data to be evaluated by a medical professional before implementing.
I can compute items to take, those computations do not provide solid information on rotations, dosages, etc.
I cannot tell people what they should take or not take. I can inform people items that have better odds of improving their microbiome as a results on numeric calculations. I am a trained experienced statistician with appropriate degrees and professional memberships. All suggestions should be reviewed by your medical professional before starting. Some suggestions may be counterindicated for other medications you are taking and medical conditions.
The answers above describe my logic and thinking and is not intended to give advice to this person or any one. Always review with your knowledgeable medical professional.








Recent Comments