This is a follow up of several prior posts
- Jul 8, 2022. A Complex Microbiome Situation
- Sep 9, 2022 Another ME/CFS Microbiome Follow Up
This person did his tests using OmbreLabs.com and then transfer the data to biomesight.com.
I did the special studies for a couple months taking symbio, florastor, GOS,Arabox., d ribose, pea fiber, etc.
Feel in general more energized, specially after the round of florastor which I had done just a four days then tested at the time so impact probably won’t fully show in this test.
Where to go from here? Drop special studies focus or stay the course?
Why Follow Up Posts are important
The first item is simple, does the model and suggestion appear to work. Everything is theoretically computed, not based on clinical practice or clinical studies. The second item is that these posts encourages people to try suggestions, or to do “self-serve” with the site.
Foreword – and Reminder
I am not a licensed medical professional and there are strict laws where I live about “appearing to practice medicine”. I am safe when it is “academic models” and I keep to the language of science, especially statistics. I am not safe when the explanations have possible overtones of advising a patient instead of presenting data to be evaluated by a medical professional before implementing.
I cannot tell people what they should take or not take. I can inform people items that appears to have better odds of improving their microbiome as a results on numeric calculations. I am a trained experienced statistician with appropriate degrees and professional memberships. All suggestions should be reviewed by your medical professional before starting.
Comparisons between Samples
We will start by adding new columns for the latest sample. The person measured with Ombre and then transferred data to Biomesight to get a second interpretation of the raw digital data.
| Criteria | BS 6/6 | BS 7/19 | BS 11/22 | OL 6/6 | O 7/19 | O 11/22 |
| Lab Read Quality | 2.1 | 5.4 | 4.2 | 2.1 | 5.4 | 4.2 |
| Bacteria Reported By Lab | 280 | 497 | 468 | 365 | 628 | 473 |
| Bacteria Over 99%ile | 2 | 7 | 0 | 5 | 6 | 1 |
| Bacteria Over 95%ile | 24 | 31 | 2 | 27 | 24 | 9 |
| Bacteria Over 90%ile | 49 | 58 | 12 | 49 | 51 | 21 |
| Bacteria Under 10%ile | 17 | 62 | 187 | 18 | 60 | 87 |
| Bacteria Under 5%ile | 5 | 30 | 90 | 10 | 28 | 46 |
| Bacteria Under 1%ile | 0 | 13 | 22 | 1 | 7 | 2 |
| Rarely Seen 1% | 0 | 4 | 0 | 0 | 9 | 0 |
| Rarely Seen 5% | 4 | 18 | 15 | 8 | 40 | 9 |
| Pathogens | 15 | 25 | 39 | 19 | 28 | 33 |
| Outside Range from JasonH | 4 | 4 | 9 | 7 | 2 | 9 |
| Outside Range from Medivere | 17 | 17 | 20 | 16 | 16 | 20 |
| Outside Range from Metagenomics | 7 | 7 | 7 | 7 | 7 | 7 |
| Outside Range from MyBioma | 9 | 9 | 10 | 14 | 14 | 10 |
| Outside Range from Nirvana/CosmosId | 22 | 22 | 22 | 23 | 23 | 22 |
| Outside Range from XenoGene | 6 | 6 | 10 | 11 | 11 | 10 |
| Outside Lab Range (+/- 1.96SD) | 6 | 13 | 2 | 10 | 14 | 2 |
| Outside Box-Plot-Whiskers | 70 | 84 | 29 | 64 | 61 | 25 |
| Outside Kaltoft-Møldrup | 70 | 113 | 70 | 112 | 182 | 91 |
| Condition Est. Over 99%ile | 1 | 1 | 0 | 0 | 0 | 0 |
| Condition Est. Over 95%ile | 2 | 4 | 0 | 0 | 0 | 0 |
| Condition Est. Over 90%ile | 5 | 6 | 0 | 2 | 2 | 0 |
| Enzymes Over 99%ile | 3 | 10 | 1 | 13 | 15 | 5 |
| Enzymes Over 95%ile | 46 | 32 | 16 | 69 | 82 | 147 |
| Enzymes Over 90%ile | 90 | 51 | 103 | 155 | 411 | 405 |
| Enzymes Under 10%ile | 102 | 219 | 354 | 55 | 138 | 169 |
| Enzymes Under 5%ile | 45 | 132 | 154 | 22 | 67 | 78 |
| Enzymes Under 1%ile | 6 | 47 | 29 | 5 | 2 | 0 |
| Compounds Over 99%ile | 9 | 7 | 29 | 104 | 126 | 19 |
| Compounds Over 95%ile | 56 | 76 | 233 | 385 | 397 | 89 |
| Compounds Over 90%ile | 292 | 313 | 347 | 533 | 548 | 118 |
| Compounds Under 10%ile | 72 | 125 | 264 | 183 | 248 | 133 |
| Compounds Under 5%ile | 39 | 64 | 163 | 109 | 127 | 100 |
| Compounds Under 1%ile | 5 | 21 | 41 | 16 | 17 | 42 |
What do I see above?
- Sample Quality are the same (expected from using the same FASTQ file)
- Rare and very high bacteria have a significant improvement
- All of the statistical out-of-range measures(Std Dev, Box-Plot, K/M) reduced the count significantly.
- Most of the expert suggested ranges increased.
- Condition profiles dropped to zero.
- We see the fragileness of some measures to the software being used to interpret the raw data.
- Enzymes Over… one dropped from prior and the other increased from the prior
My general opinion is that the person has improved objectively. The algorithm explicit goal is to reduce all of the statistical out-of-range measures. Ideally, it will also “fix” the person but that is more complex, we lack sufficient knowledge to hand pick the bacteria. We can get associations to specific bacteria — association is NOT causality often (despite many politicians claiming such!).
Going Forward
KEGG Computed Probiotics
Both labs resulted in the same priority: Escherichia coli at the top, the soil based mixtures, then Bacillus subtilis (I personally prefer to get it “au natural”, i.e. in the traditional Japanese Soy based food, Natto).

Special Studies Numbers
I am going to skip them, mainly because the results are erratic until I get a better understanding of this. See Caution: Special Studies Suggestions.
There is a possibility of both being right. Right meaning shifting from the current dysfunctional equilibrium. This could be visualize as shown below. I am seeking understanding and building different approaches. Each approach could work for some (but not others). Too many factors for certainity.
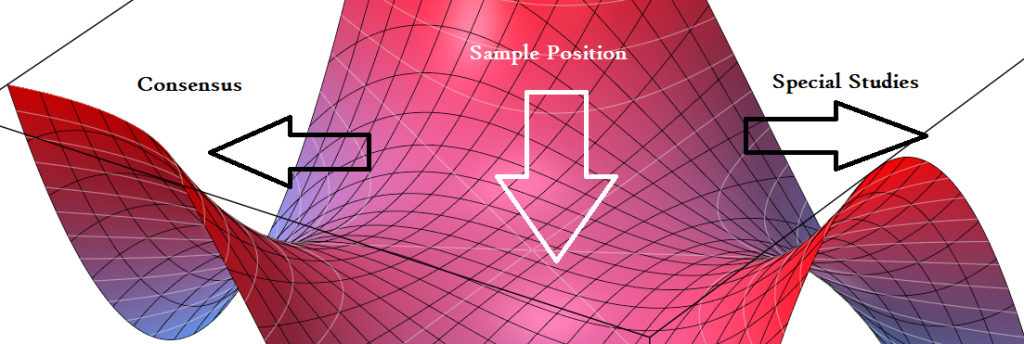
Going forward
I am building a consensus report from the items marked 2 above using the Special Studies. The list is similar to other people with ME/CFS. We see 2 E.Coli probiotics (symbioflor 2 e.coli probiotics, colinfant e.coli probiotics) at the top with d-ribose (a sugar used by E.Coli). This is then followed by the earth based probiotics( General Biotics Equilibrium, Prescript Assist (Original Formula), Prescript Assist (2018 Formula)).
For Consensus building, I am going to use the three statistical selectors with both Ombre and BiomeSight:
Then we will do an uber-consensus (combining the consensus from each)
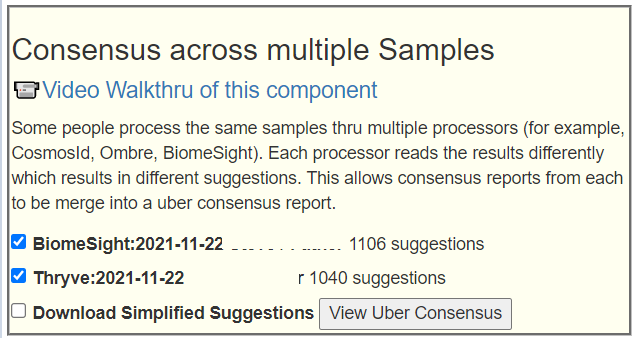

The suggestions look very typical for ME/CFS persons. I attached the Simplified Suggestions
The probiotics suggested are
- akkermansia muciniphila
- bacillus clausii
- bifidobacterium (animalis)lactis
- bifidobacterium breve
- enterococcus faecium
lactobacillus brevislactobacillus bulgaricuslactobacillus caseilactobacillus paracaseilactobacillus rhamnosuslactobacillus salivarius- pediococcus acidilactici
I have marked the lactobacillus to indicate that they do not play well with E.Coli probiotics. Thus, should be in an alternative probiotic cycle.
We see by individual fibers, that the separate suggestion of a low fiber diet

The New Boy on the Block: Food Suggestions
See New Suggestions Page — Food Centric. This was trying to mine the available data more to get practical suggestions.
- Flavonoids: Only Barley was a positive (for Ombre) , it was not on Biomesight list
- Food Contents: Again, only Ombre had positive suggestions: Brazil Nuts and Olive Oil. Biomesight data disagreed on the Brazil Nuts
Why so few? Why labs contradict? This comes down to two key challenges: Different interpretation of the bacteria from the digital data; a low volume of studies on the substances we are using to build food suggestions. Also, the suggestions are based on some of the contents of the food; there are other parts of the food that will have other effects. This is why direct food, herb and spice studies are best. Every food is a complex mixture of chemicals. Some may help, some may hurt. Care must be taken to avoid the simplistic logic that “Super Breakfast Food contains barley, thus it is good/healthy to eat!” Ignoring the 10 grams of sugar in this product.
Thus, these suggestions should be taken with a grain of salt. They are better than random choices, but far short of what we would ideally like.
Bottom Line
I ran some of the Special Studies suggestions and did a download of simplified consensus. Between approaches, we had agreement on taking:
- Probiotics:
- akkermansia muciniphila
- bifidobacterium (animalis)lactis
- lactobacillus salivarius
- saccharomyces boulardii
- Other
- Vitamin K2
- Calcium
- Echinacea
- Omega-3 fatty acids
- Pomegranate
- Rutin
- Tea Tree oil
As well as agreement on avoiding
- fructooligosaccharides (FOS)
- jerusalem artichoke
- Flaxseed
- Vitamin B2 Riboflavin
“This is too complicated” is what I can hear some people saying. This analysis digs into the nature of the data which is really not needed for most people. I am trying to get better understanding. It looks at some of alternative methods of getting suggestions. It is likely of interest to those treating microbiome dysfunctions as it illustrates many of the challenges in interpreting.
For most people, the best process stays the same:
- Upload the data
- Try several different ways of generating suggestions, building a consensus from
- Look at the consensus
Why is consensus important? Simple, we have very incomplete data and also have limited accuracy with the microbiome tests. Going the consensus approach is similar to using a Monte Carlo Simulation, an appropriate approach to deal with complex processes with many parameters that are fuzzy that produces better results.
Recent Comments