This post came out of a Facebook dialog with someone reporting they are taking Symbioflor-2 and not seeing any changes in test results.
This study says it well: “16S rRNA sequencing is not diagnostic for Enterobacteriaceae or Escherichia/Shigella“. Rethinking gut microbiome residency and the Enterobacteriaceae in healthy human adults, 2019
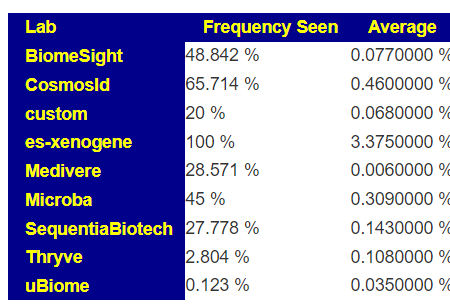
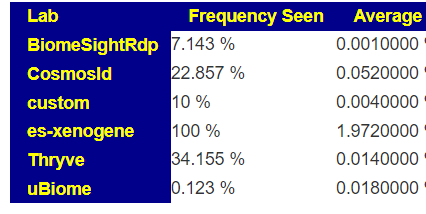
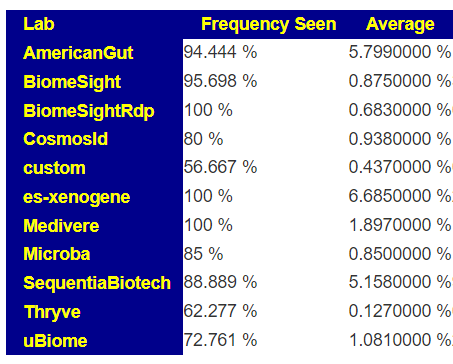
The numbers reported on most tests for these bacteria are extremely questionable. The one exception is Xenogene (based in Spain). This can be seen on these summary pages. These bacteria are grossly under reported on 16s tests.
What does this mean for manipulation? If you take Symbioflor-2 or Mutaflor, you may not see any changes in your tests (or they may become worse), when in reality they have taken up residency and are increasing. Either you go and do tests with Xenogene; or you use subjective measurements. My subjective measurement from Mutaflor was a massive severe herx for the first two weeks.
Some other recommended readings:
Recent Comments