I have done open sharing of data at http://lassesen.com/ubiome in formats designed for data science import and processing. In this post, I taken some of this data and make it uploadable to our new database schema. I will also work on utilities to allow data to be exported and shared going forward. If you get data donated on your site (AND you obtain consent for sharing the data anonymously) remember to share it with other citizen scientists.
There are a series of tables that needed to be exported and then imported. The contents of the export is illustrated below. Note: Guid in the export will be converted to an Id by the import.
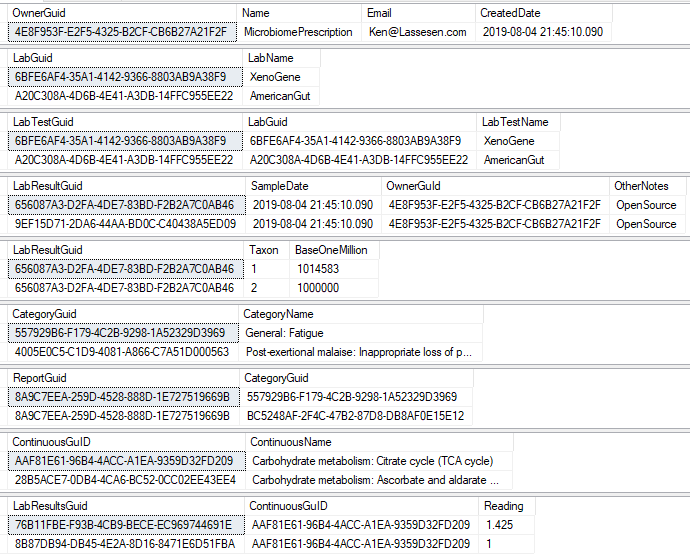
To deal with some issues (and to support interchange of data between people using this open source, I added two GUID columns. This means that every sample uploaded to a site, or report given to a site is unique and may be exchange between sites without duplicates. If the same microbiome report is uploaded to multiple sites, then there will be duplicates — until you de-duplicate the data (see this earlier post).
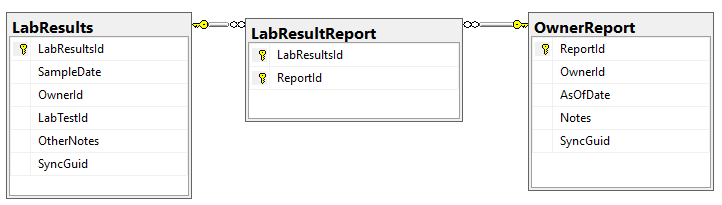
The process is simple:
- Bring in all of the reference data (Continuous, Category, Labs, LabTests definitions) using their names as keys to prevent duplication
- Import the lab and other results — using SyncGuid to prevent duplication.
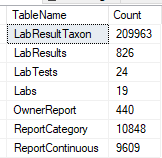
This is done by passing the XML from one site to a stored procedure (using Datasets) and then processing it. It is important to test the code to insure that the same data being imported multiple times does not result in extra rows. The TSQL below gives the counts on key tables.
Select 'LabResultTaxon' as TableName, Count(1) as Count from LabResultTaxon UNION Select 'LabResults' as TableName, Count(1) as Count from LabResults UNION Select 'LabTests' as TableName, Count(1) as Count from LabTests UNION Select 'Labs' as TableName, Count(1) as Count from Labs UNION Select 'OwnerReport' as TableName, Count(1) as Count from OwnerReport UNION Select 'ReportCategory' as TableName, Count(1) as Count from ReportCategory UNION Select 'ReportContinuous' as TableName, Count(1) as Count from ReportContinuous
After the import, you should recalculate core statistics for each LabTest type (see this post for a reminder). Every thing has been rolled into the Library, so the program is very short (but the library code and tSQL code is not).
var import = new DataSet(); import.ReadXmlSchema(schemaFile.FullName); import.ReadXml(exportFile.FullName); DataInterfaces.ConnectionString = "Server=LAPTOP-...."; DataInterfaces.Import(import); LabTests.ComputeAllLabs(4);
Source for Importer: https://github.com/Lassesen/Microbiome2/tree/master/Import
Data Location: https://github.com/Lassesen/Microbiome2/tree/master/Import/Data
Updated SQL Server Schema: https://github.com/Lassesen/Microbiome2/tree/master/DBV2
1 thought on “Populating with Open Data”
Comments are closed.