Background
- A nasal swab showed large amounts of staph aureas (but no MARCoNS) and k. oxytoca.
- A stool test from Doctor’s Data that was done prior to SIBO treatment and treatment with a rotation of probiotics and prebiotics.
- Partially hydrolyzed guar gum (PHGG) gave me a lot of bowel mucus after a few weeks, so I stopped it.
- Tested with NirvanaBiome
Analysis
Because of the mention of SIBO and because I had just finished refactoring Symptom to Bacteria association, I decided to start there. I do not have great expectations because published clinical studies were unable to find any significant patterns.
See Symptoms Associations post for how we are doing this
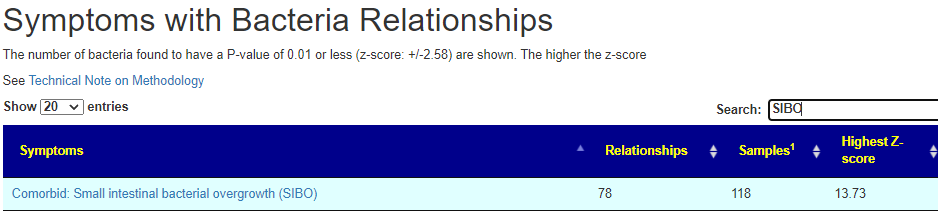
We now see hoe well this person matches..
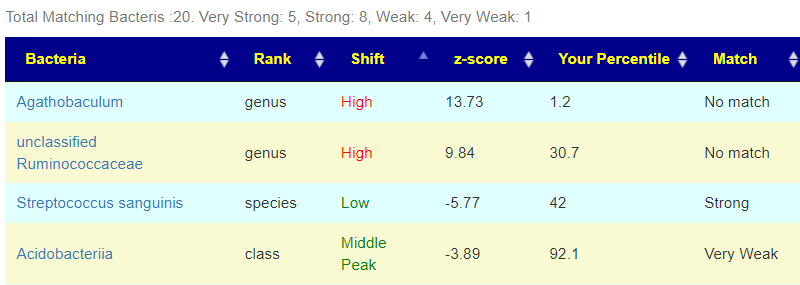
Looking at Other Explorers
- KEGG Enzymes – nothing
- KEGG Module:33. Very Strong: 29, Strong: 4 – this is interesting… especially since all 33 are Middle Peaks. This should be explore in time. For anyone interested in exploring,
- beta-Oxidation, acyl-CoA synthesis
- Biotin biosynthesis, pimeloyl-ACP/CoA => biotin
- CAM (Crassulacean acid metabolism), light
- Citrate cycle (TCA cycle, Krebs cycle)
- Citrate cycle, second carbon oxidation, 2-oxoglutarate => oxaloacetate
- CMP-KDO biosynthesis
- Cobalamin biosynthesis, cobyrinate a,c-diamide => cobalamin
- D-Galacturonate degradation (bacteria), D-galacturonate => pyruvate + D-glyceraldehyde 3P
- Gluconeogenesis, oxaloacetate => fructose-6P
- Glycogen degradation, glycogen => glucose-6P
- Glycolysis (Embden-Meyerhof pathway), glucose => pyruvate
- Glycolysis, core module involving three-carbon compounds
- Histidine biosynthesis, PRPP => histidine
- Histidine degradation, histidine => N-formiminoglutamate => glutamate
- Inosine monophosphate biosynthesis, PRPP + glutamine => IMP
- Isoleucine biosynthesis, pyruvate => 2-oxobutanoate
- Leucine biosynthesis, 2-oxoisovalerate => 2-oxoisocaproate
- Lysine biosynthesis, DAP dehydrogenase pathway, aspartate => lysine
- NAD biosynthesis, aspartate => quinolinate => NAD
- Pantothenate biosynthesis, valine/L-aspartate => pantothenate
- Pentose phosphate pathway (Pentose phosphate cycle)
- Pentose phosphate pathway, non-oxidative phase, fructose 6P => ribose 5P
- Pentose phosphate pathway, oxidative phase, glucose 6P => ribulose 5P
- Phosphate acetyltransferase-acetate kinase pathway, acetyl-CoA => acetate
- Phosphatidylethanolamine (PE) biosynthesis, PA => PS => PE
- Pimeloyl-ACP biosynthesis, BioC-BioH pathway, malonyl-ACP => pimeloyl-ACP
- Pyrimidine ribonucleotide biosynthesis, UMP => UDP/UTP,CDP/CTP
- Riboflavin biosynthesis, plants and bacteria, GTP => riboflavin/FMN/FAD
- Serine biosynthesis, glycerate-3P => serine
- Tetrahydrofolate biosynthesis, GTP => THF
- Tryptophan biosynthesis, chorismate => tryptophan
- Uridine monophosphate biosynthesis, glutamine (+ PRPP) => UMP
- Valine/isoleucine biosynthesis, pyruvate => valine / 2-oxobutanoate => isoleucine
- KEGG Product:50. Very Strong: 36, Strong: 5, Weak: 6, Very Weak: 2
Adjusting for Middle Peak
Finding middle peaks presents some challenges for suggestions. The graphics below may help explain with a middle peak is. The why is not simple, but likely a complex interaction of many things (like other bacteria)
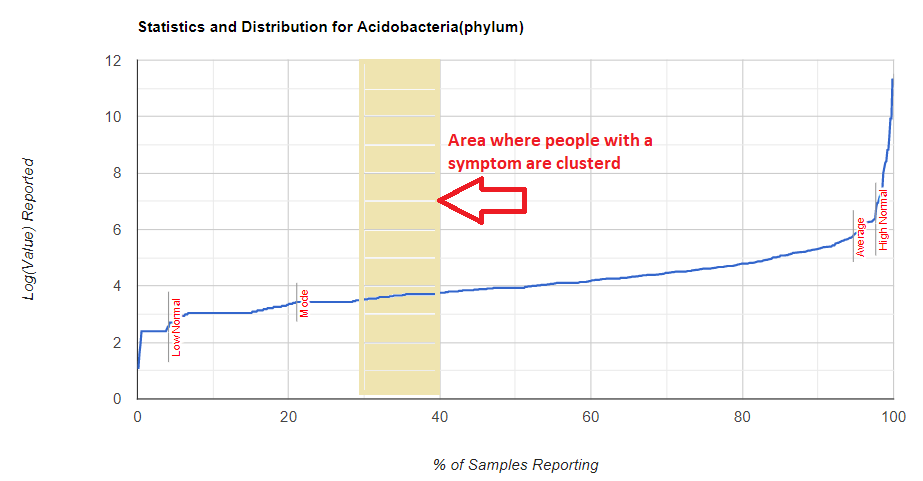
But looking at the raw numbers will have most people seeing “nothing”
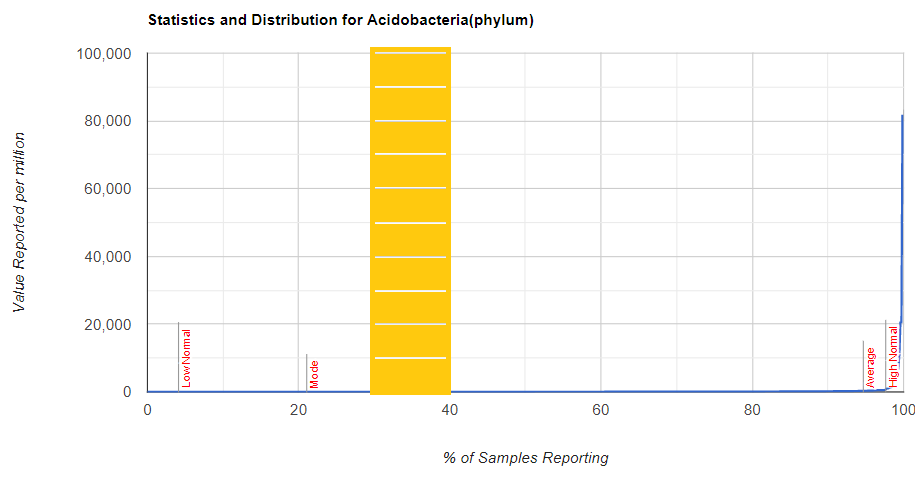
As a result of doing this post, I realized that it was possible to generate suggestions for these middle peaks. This is explained in the video below
The resulting suggestions are below. These are very blinkered suggestions that ignore extreme values.
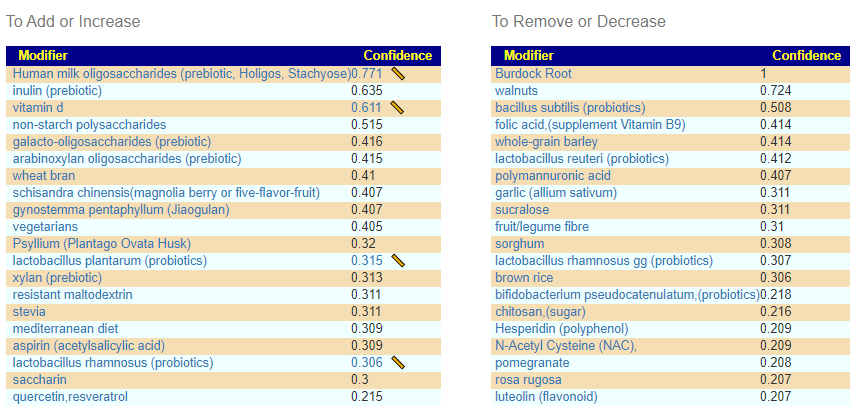
Consensus Suggestions
See Multiple Conditions Microbiome Analysis post for more background.

I then did two common suggetions
- Kaltoft-Moltrup Suggestions
- Jason Hawrelak
This results in 3 items in the Consensus
KEGG Suggestions
The clear winner is Sun Wave Pharma/Bio Sun Instant probiotics – consisting of Clostridium butyricum and Bacillus mesentericus.
For supplements, we have
- beta-alanine
- iron
- L-Phenylalanine
- L-Tyrosine
- Vitamin B-12
Medical Conditions(PubMed) had only one item that was border line item, Graves’ disease (an immune system disorder that results in the overproduction of thyroid hormones (hyperthyroidism). A simple blood test should clarify this risk.
Thanks very much – Based on a quick skim, it’s interesting that Graves comes up bc my dad has it and my TSH has been declining the last couple of times it was measured, but they didn’t check my T4, so I need to have another test soon that will look at actual hormone levels.
From person after reviewing the draft,
Putting it all together – Consensus
On my first drafts of this post, I did the manually and then realized automating it was a much better solution for me and for others.
REMINDER: The numbers are NOT the degree of impact. It is the confidence that it will help. That is, the number of bacteria shifted in the right direction, the number of studies finding the shifts and the amount of shifts we want to see.
:
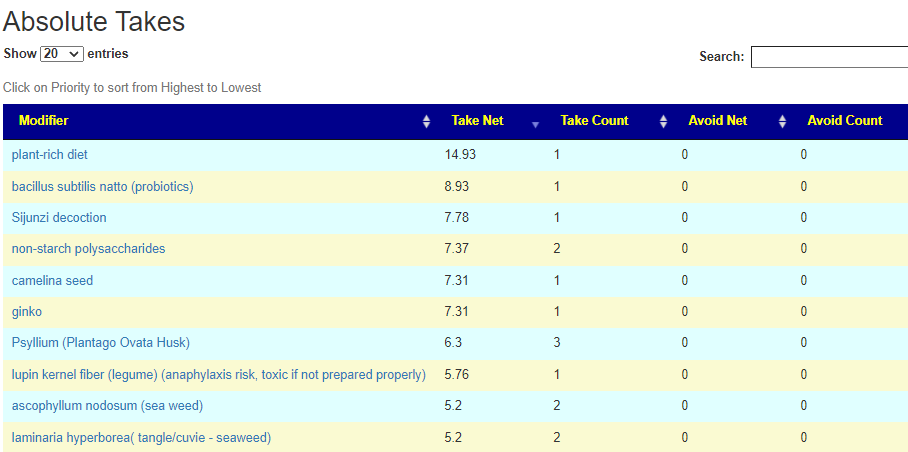
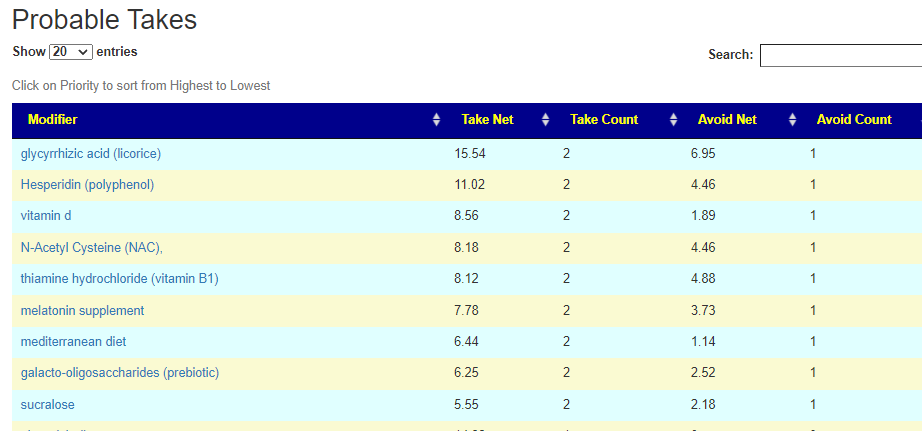
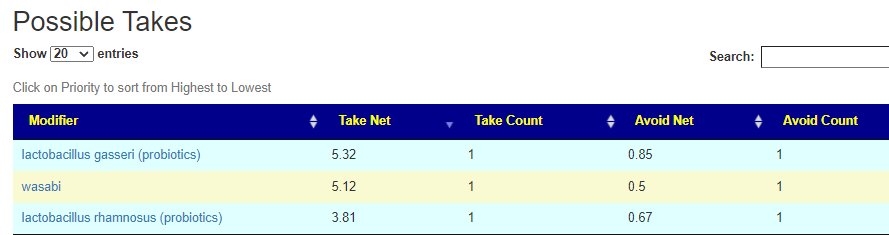
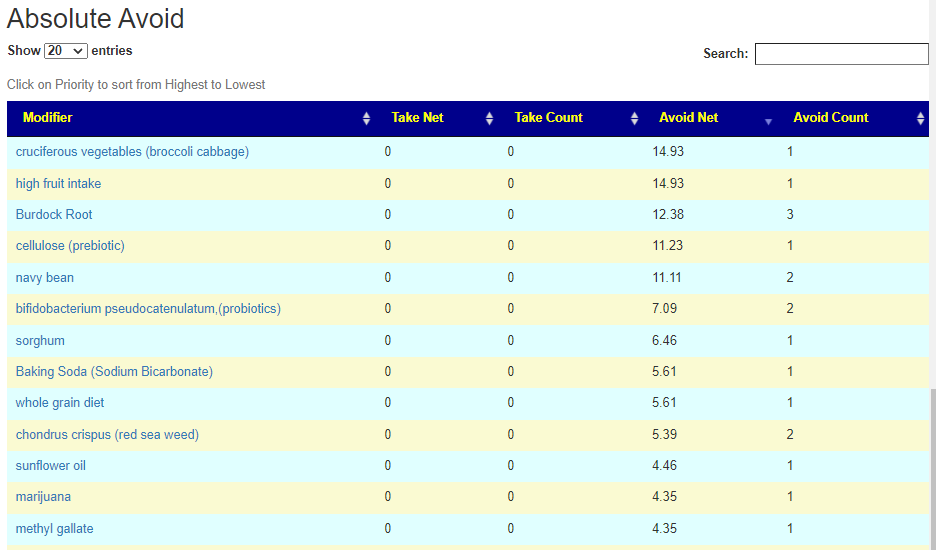
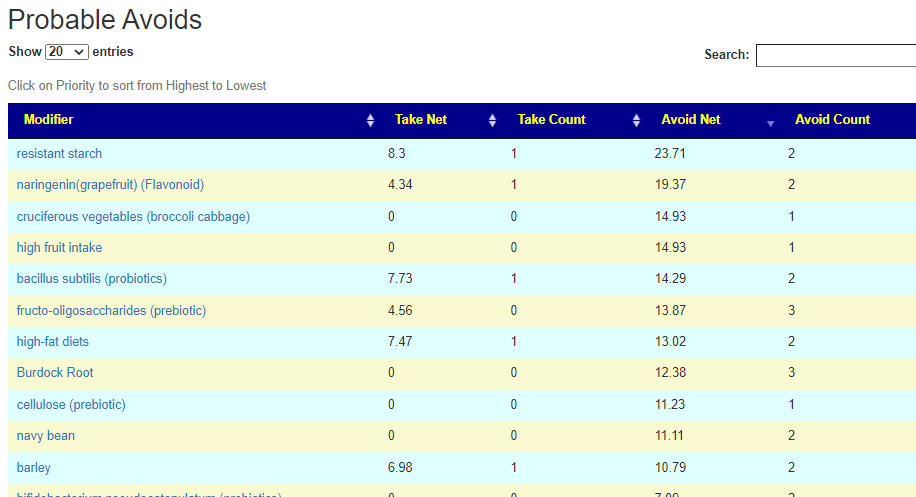
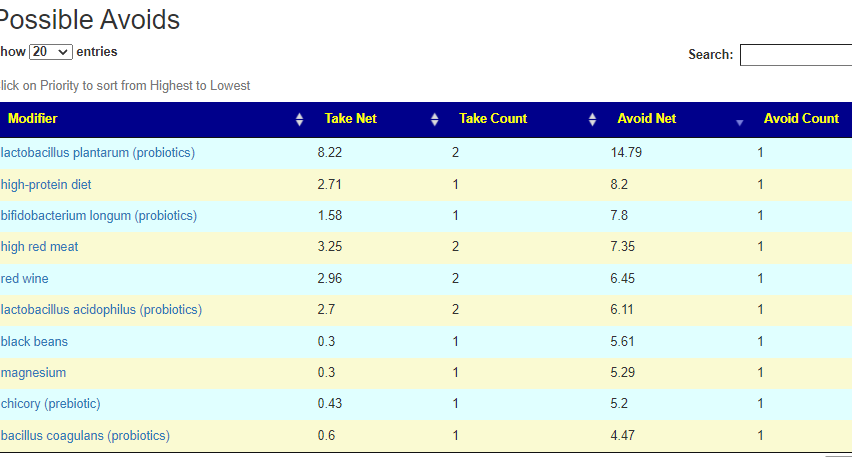
There is now a better way!
I have enabled a Consensus Report on all suggestions done on a specific sample in the last 24 hours. After 24 hours, I remove the data, First, I will do the two most common suggestions:

After the 2nd one, a new choice appears on the home page sample menu as shown below
I then added Comorbid: Small intestinal bacterial overgrowth (SIBO) to the mixture,
I ahd tried a fourth one, I went to Advance Suggestions and selected PubMed for SIBO
Now, let us look at the Consensus View, it’s divided into 6 sections as shown below
NOTE: by default it is sorted by name. Double click the value column and then double click the Count column will sort it as shown below (i.e. best or worst at top of list).
Recap
This was an interesting analysis in that PubMed literature failed to find any matches, while citizen science found patterns. The patterns were often middle peaks which would not have been seen with cookbook statistical methods focused on normal distributions (bell curves)
Suggestions should be review by your medical professional. All of the items are based solely on the impact on the microbiome bacteria — often items may have adverse effect on medical conditions.
One of the confusing aspects of Microbiome Prescription is that suggestions are derived through different paths with suggestions from different paths being in partial contradiction. Each path has limited and often fuzzy data. The safest path is to merge the lists with what is in common, the next step forward (if the first step does not give you too much to do), is items that is cited in one list and not contradicted in another list.
Recent Comments