For people amusement, while reviewing new studies on PubMed, I got a pleasant surprise! My citizen science project (MicrobiomePrescription) is referenced in BMC Bioinformatics article published June 3 2021. Adding to my prior mentions in New Scientist and The Lancet. “Dysbiosis of the gut microbiome is a risk factor for osteoarthritis in older female adults: a case control study“
Most errors will automatically send emails about the issue and they should be resolved within 24 hrs. If something is still broken after 24 hours, please email Research@MicrobiomePrescription.com.
Currently the online database consists of 50 million rows of data.
Caveat Emptor
This is a “best effort” project. Data may be incorrectly interpreted or entered on occasion. It is believed that the entry accuracy is between 95% and 98%. Links to the source study are provided for people to verify information before relying on it. There is no claim of “correctness”, just reasonable efforts.
2024
- June 20th, added High Histamine producers to samples and explicit 🔨 reduce all bacteria associated.


- June 18th added report across all pub med studies flagging bacteria association for different conditions
- June 14th release MD centric report with a few conditions in 8 languages.
- June 12th Added computation of biofilm forming bacteria
- June 4th, refactor compensate for antibiotic use with a sample. Focused on the lowest percentiles ones that are impacted by the antibiotic.
- May 26th, removed MTHFR from analysis. PrecisionBiome.Eu during their audit of the information, identified that the wrong KEGG enzyme was being cited -typing error (1.5.1.5 instead of 1.5.1.53). Insufficient data to warrant use.
- May 23rd, added suggestions for symptoms across all citizen science findings. See bottom of this post.
- May 20th, revised PDF report by adding this section

- May 18th, Added ability to hide Prescription Items on Consensus
- May 17th, Update algorithm for just give me suggestions, see _Algorithm for “Just Give Me Suggestions”
- May 15th, Added Critical Bacteria as a filter, see Technical Note: Identifying Key Bacteria to Address
- May 2nd, add Chuckling Goat transcription form
- May 1st, released localized names for microbiome modifiers in English, German, French, Italian, Spanish, Danish, Swedish and Welsh. Thanks to PrecisionBiome.eu
- April 30th, Vitract team of post graduate employees indicated that they have completed most of their reviews of microbiome modifiers to bacteria. A few minor corrections were made.
- April 27th, added bacteria to bacteria impacts, see Technical Note: Bacteria Influencing Bacteria
- April 15-30th, evaluate comments from Vitract‘s team of post graduates in Microbiology who are reviewing citations.
- April 10th, Fixed some suggestions bugs with advanced methods. Some were bugs with newer samples only (changes of methodology for computing percentiles)
- Hand Picked reworked and clarification added to the page
- Increase Butyrate
- Only Use Bacteria deemed Unhealthy
- Increase Anti-Inflammatory Bacteria
- April 8th, Released Enzymes reworked with expanded description
- April 6th, Released Compounds reworked with expanded description
- April 5th, Connects Modifier summaries to Suggestions and Consensus pages
- April 4th, started building out Modifier summaries using ChatGPT for initial data. https://microbiomeprescription.com/Library/ModifierTable
- April 2nd, issue with KM Suggestions reported. Some code was missed in updating sample percentiles (old samples worked, new samples did not)
- March 31, resolve issue with Thorne results not displaying on the Taxon Tree.
- March 30, Resolved inconsistency in Computing PubMed Conditions. Added Charts for PubMed Conditions on Health Analysis and update charts links to current.
- March 29 Walkthru of new and revised features on the Health Indicators Page (youtube.com)
- Audited, found errors and corrected items on Health Indicators Pages
- Added charts for measures on that page.
- March 25, Expanded and corrected External Reference Ranges for different bacteria (Lookup Bacteria page)
- March 24, Added GABA producers to Probiotic Mixtures.
- March 20, Updated Individual Suggestions to use refactored Taxon Percentiles (had been missed in refactor) https://microbiomeprescription.com/library/gutmodifiers?sampleId=….
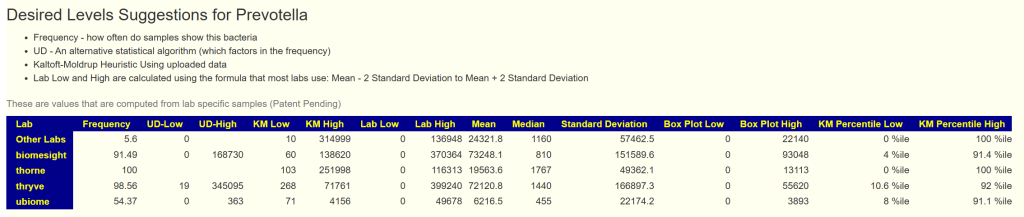
- March 18, add new experimental reference range (UD-Low and UD-High
- March 4-18: Down with COVID — bugs may occur. Please report as you find them
- March 13: User identified processing issues with data from Extensive impact of non-antibiotic drugs on human gut bacteria. All old data (over 100,000 facts) deleted and data re-entered.
- The relative impact of each was incorporated in the revised fact.
- Some new drugs, substances were added, for example Piracetam, the nootropic.
- Some suggestions are expected to shift.
- March 12: Add Mold Index to site, fixed Symptom forecasts to be out of 100.
- March 11: Restored Adding Symptoms functionality. Broke by a refactoring elsewhere
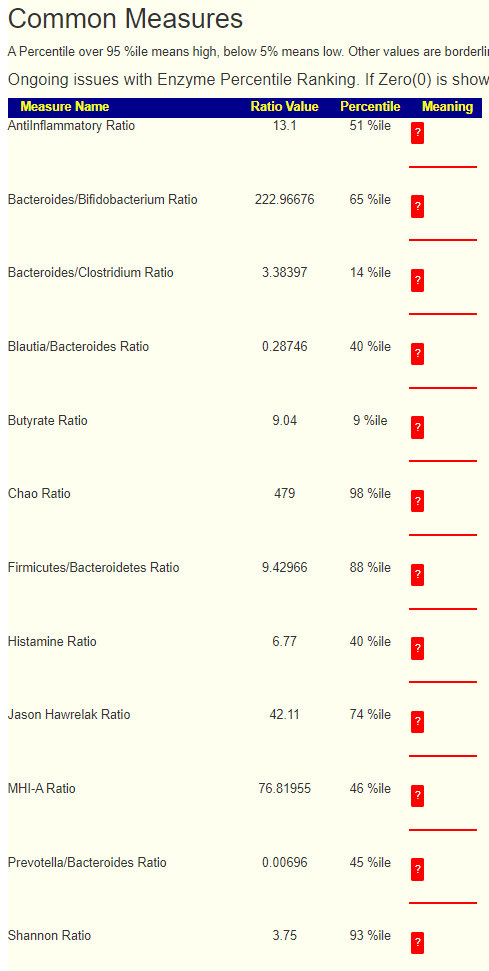
- Match 10: Refactor Common Measure to use lab specific data for percentile, updated the ? Meaning for missing items
- March 9 Taxa Association to symptoms expanded to allow filtering by different taxa ranks (not just genus). For each option, a priori suggestions were added.
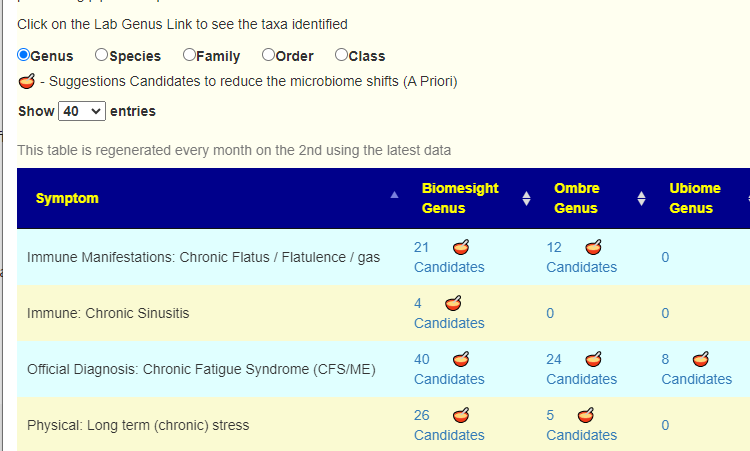
- March 2 – Monthly update of Percentile and other regressions scheduled to occur on the 2nd of each month, Updates will usually cause only minor shifts of suggestions unless rare bacteria is involved, then more dramatic changes may occur.
- Feb 28 – Refactor Genus Associations for faster execution and expanded to include uBiome and Thorne in the regression.
- Feb 26 – Refactor PubMed Medical Conditions providing more data for evaluation and a separate page/panel
- Feb 22 – Deleted a citation (on Cadium) and it’s related facts because the study link was very wrong and the correct source could not identified. One of the backup links went to a different article, that is now annotated Retracted article. Some changes of suggestions could occur as a result.
- Feb 21 – Audited Taxa Percentile and found that 1.5% were computed too low. This occurred on rare taxa.
- Feb 16 – Added Absolutely safe suggestions filter to consensus view
- Feb 14 – Added and tested two ways of selecting bacteria based on symptoms
Video on above 2 features: https://www.youtube.com/watch?v=CdrRCyQNzfs
- Feb 9th, User pointed out an error in a few Bacteria description (wrong bacteria described). Implemented audit and correction of 1,336 descriptions.
- Feb 7th, deprecated old symptom forecasting method, replacing with new genus based algorithm.
- Feb 6th, Investigated issue with two uploads of the same data reporting different results. Explanation:
When an upload happens, I create missing hierarchical levels by summing up the children. This means that two identical samples uploaded at different times may different missing levels if the Hierarchy table was updated or modified between samples. - Feb 6th, Refactor the PubMed studies to support selecting multiple conditions concurrently. 16% is a toss up, below 16% is unlikely to have a condition. The higher the percentage match, the more likely.
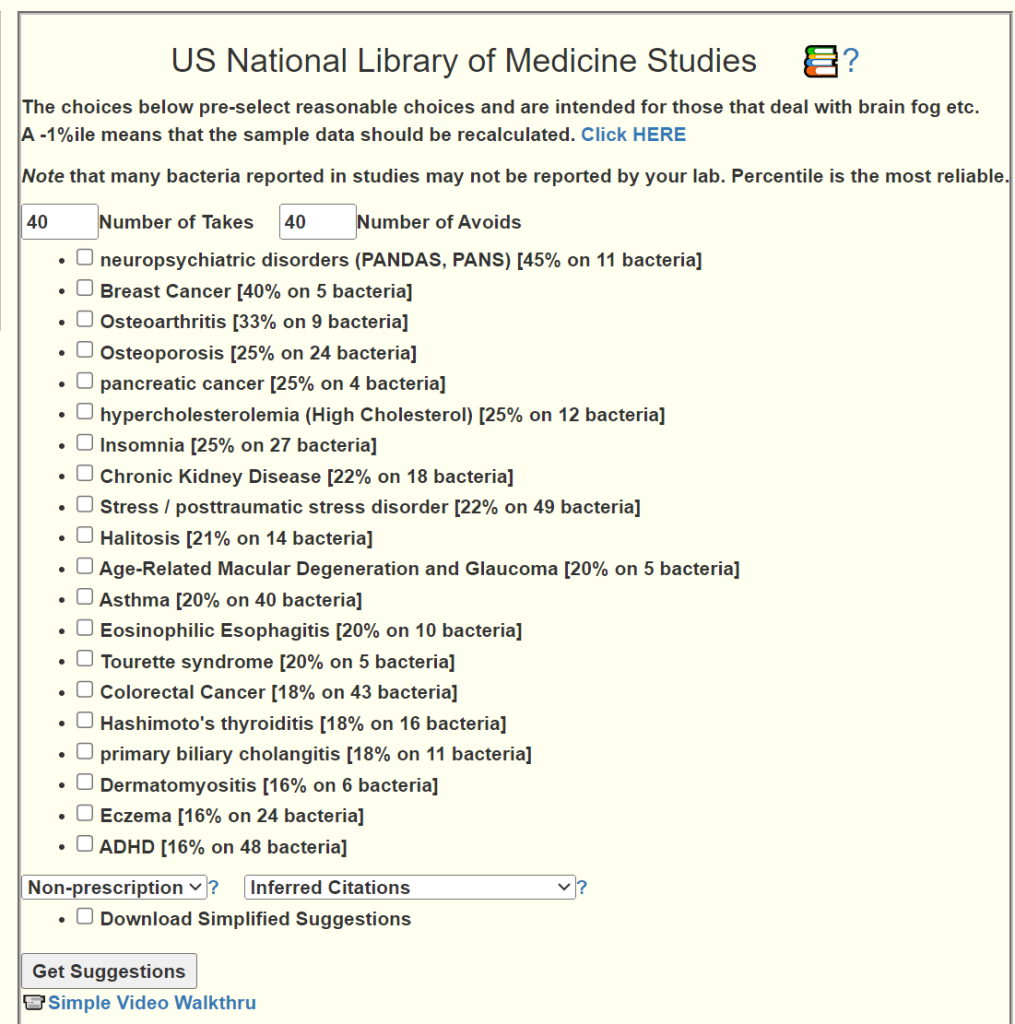
- Feb 4 — Finished refactoring of Percentiles on Enzymes, Compound and Taxons. The refactor reduced the database size by over 50% and should eliminated most stale data. There may be some bugs as side-effect. Please report them ASAP.
- Feb 3 Remove this obsolete section. It was an early punt prior to getting consensus working well. It used only one method and as a result, it was often in disagreement with the consensus.
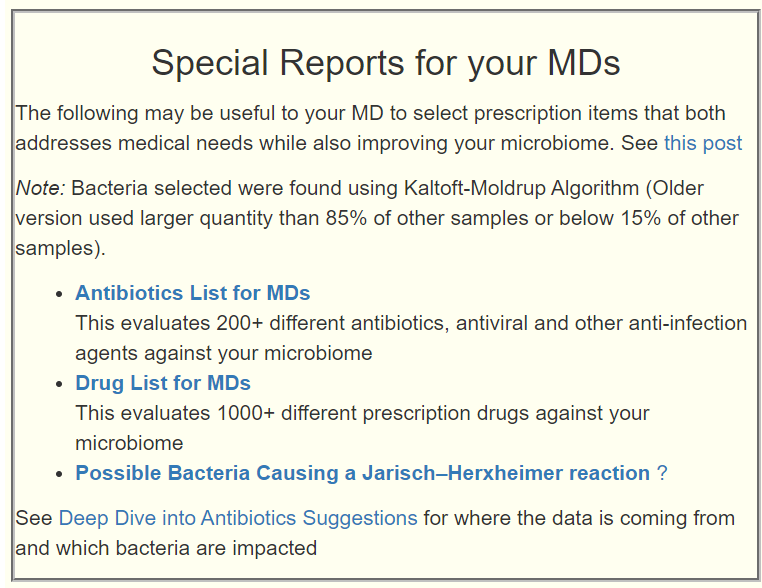
- Feb 2 Revised KEGG section to allow individual suggestions derived from trying to tune using Kegg Compounds. See Technical Demo: Using Compounds and Enzymes to get Suggestions (youtube.com)
- Feb 1 – Satisfied Jan 25th bug has been fixed
- Jan 30 — Removed the old metabolite section. It had been marked obsolete for over a year and it was time to remove it after confirming that it was likely giving incomplete information. See Technical Notes: Balancing Priorities for Fixing the Gut
- Jan 26, removed quasi duplicate citations. For some studies both the parent and the child are mentioned as shifting. We infer parent and descendants using each bacteria, this resulted in TWO records for some bacteria — the direct citation and a second citation thru a child bacteria. We identifies some 13,000 such records (in over 2.5 million) and removed these secondary records.
- Jan 25, Significant Bug found. It may have some impact on suggestions (bug impacted just 1 of 4 methods) At one spot, I have percentiles as 0-100 in the sample; but in reference values I used 0.0-1.0. This impacts KM only. Suggestions may change slightly
- Jan 25, Added Taxa Rank to Bacteria Details page
- Jan 24, Added Citizen Science symptoms to Bacteria Details page
- Jan 20, Added more lab ranges to database.
- Jan 17, Added TeleTest as a supported lab for transcription.
- Jan 11, fixed issues of citations link not working on Consensus View
- Jan 9th, fixed issue on https://microbiomeprescription.com/Sample/DoubleSignificant when there is a long list of parameters
- Fixed issue with Update button and then refactor so percentiles are updated with live data (ignoring duplicate uploads). Recent refactors allow that to be more efficiently done in a reasonable length of time.
- Taxon
- Enzymes
- Compound
- Complete Update on ALL uploaded samples is done once a week.
Recent Comments