These are often asked for with people often running off old research. Often the research is based on seeing an increase of one of these when a specific bacteria increase… hence. this bacteria must be producing it! This was the best that we could do from old, last millennium technology and lab tests. We have enter a new world that is summarized on Kyoto Encyclopedia of Genes and Genomes. Instead of measuring substances (never with a pure culture of a specific bacteria), we can look at the gene in each bacteria and what they produce. In other words, we can get more accurate information that is truly bacteria specific.
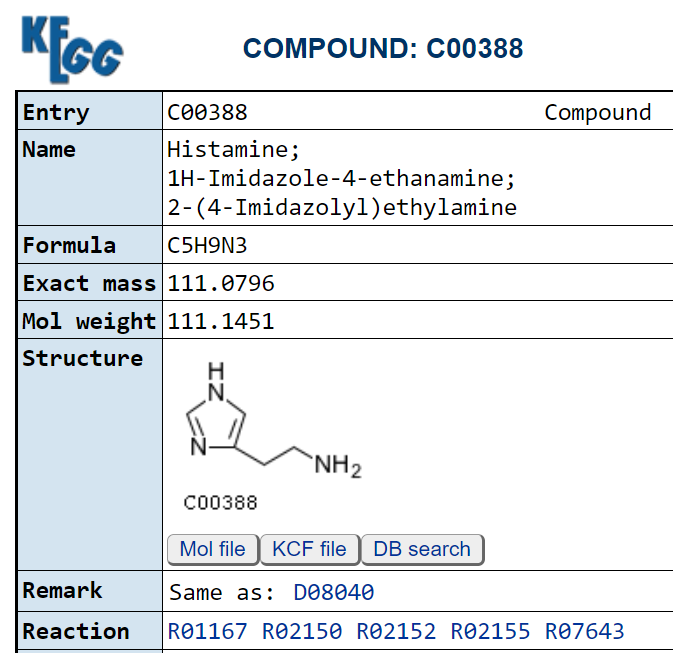
These pages list the enzymes that produces it, for example 1.4.3.22 2.1.1.8 2.3.1.- 4.1.1.22
6.3.2.18. We can then look to see if the enzyme consumes (substrate) or produces it.
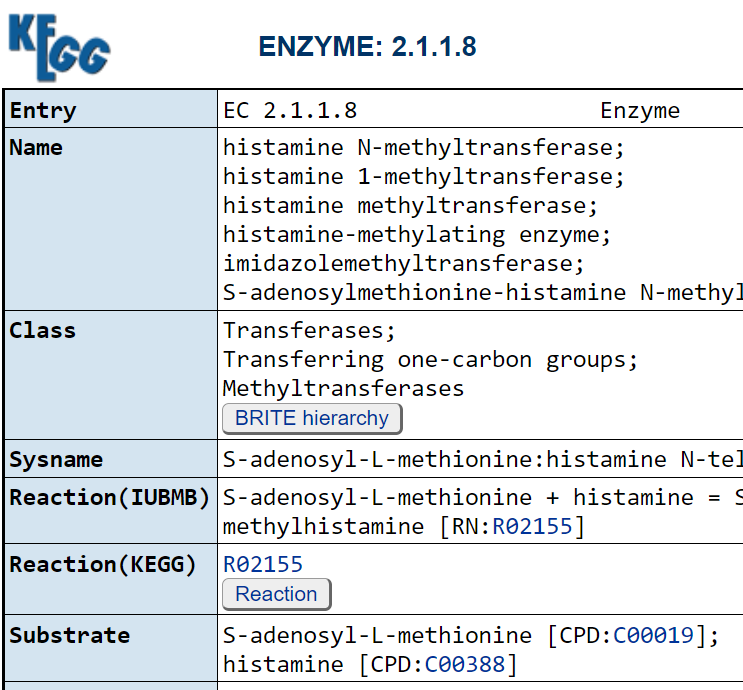
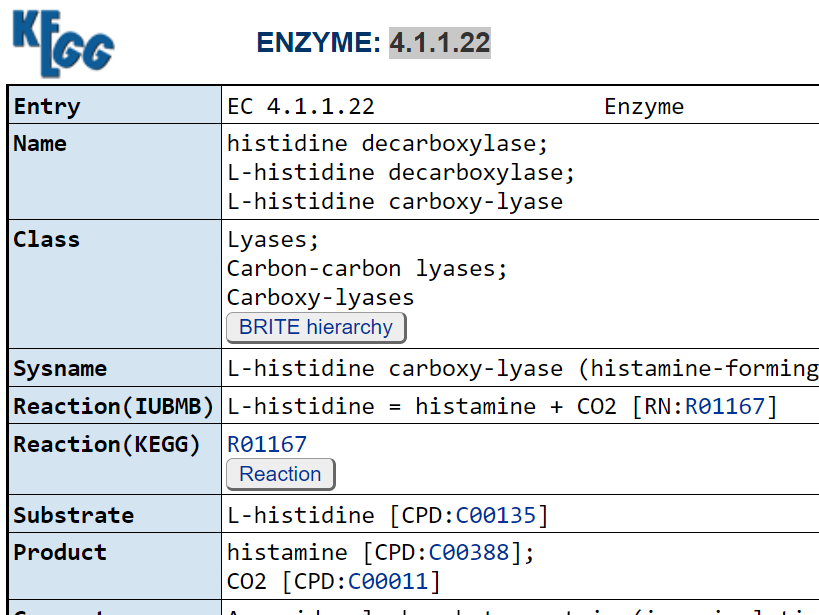
Then it is just a matter of looking up which bacteria has these enzymes!
Specific Species and Strains
The three files below are directly from KEGG data
Genus of the producers
Unfortunately, no lab detects all of these strains and often report “unknown Species in Some Genus“. I have seen that being as high as 45% not being identified in some samples. A work around is to look at the genus numbers — which are usually pretty complete for most labs. This is a two sided coin, because some members of a genus may be a producer and some may not be. We do not know the ratio for a genus between these two. This approach will identify potential candidates, but not definite candidates… more fuzzy logic.
Bottom Line
All of this information is freely available on Kyoto Encyclopedia of Genes and Genomes, it just take diligence to extract and assemble it. If you see items on a labs reporting that is NOT on these lists, challenge the lab to produce evidence. I suspect most labs will produce studies with circumstantial evidence only, “because crime goes up when this ethnic group moves in — it must be that ethnic group” — ignoring that a different ethnic group may be behind it. An example of a drug problem in one city at one time. Most of the drug dealers were African-Americans, but the importers were Vietnamese. With KEGG, we are looking for the fingerprints on the gun or drug packages intercepted,
Recent Comments