This blog is using reports from NirvanaBiome for three members of a family. NirvanaBiome use CosmosID for processing.
In this analysis we have people from the same family, implies DNA and inherited microbiome are similar.
- A 10 y.o. son who has ASD/ADHD.
- Eats various meats, dairy, grains, some beans, and a limited variety of vegetables and fruit.
- A 7 y.o. daughter who has been battling constipation for a little over a year now and had some sort of traveler’s diarrhea back in 2017 from which her GI system never seemed to fully recover IMO.
- She’s a very picky and self-limited eater. Refuses to eat any meat, but she does consume dairy in the form of yogurt and cheese and ice cream. Most of her diet is some form of processed carb, and the only fruits and vegetables she consumes are in the form of vegetable powders – potato starch based vegetable chips, granola bars with vegetable powder in them, fruit and vegetable powder added to pancakes, etc. She will eat fruit pouches that contain peach, apricot and banana, and she will sometimes eat fresh broccoli, cauliflower, some hummus and red lentil pasta.. She will drink a whey protein drink , and mother adds pea protein powder to things like pancakes as well.
- The mother diet is similar to the son’s diet
Analysis
Before getting to suggestions, I want to use the available information to understand better the three microbiome, especially since two are children. Children have a different microbiome than adults. Caution has to be taken with any data of children — we do not know what “normal” is because studies are rarely age specific.
- Aging progression of human gut microbiota [2019]
- The microbiome: An emerging key player in aging and longevity [2020]
- Unique gut microbiome patterns linked to healthy aging, increased longevity [2021]
- “We show that disease-microbiome associations display specific age-centric trends.” [2020]
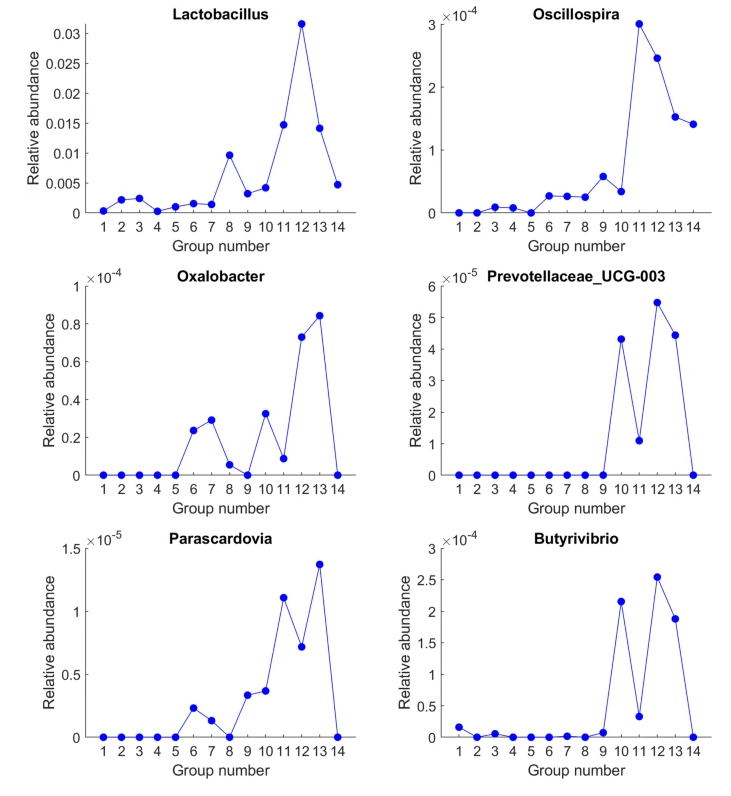
I first look at clustering of bacteria, looking at bacteria in common to all three using percentiles. The number of bacteria that had readings very close to each other was shockingly high, far more than I had expected.
| Bacteria Name | Rank | Son | Mother | Daughter |
| Ruminococcus lactaris | species | 92.4 | 93.7 | 93.5 |
| Oscillibacter ruminantium | species | 81.5 | 83.6 | 83.3 |
| Alistipes indistinctus | species | 79.5 | 76.8 | 80.2 |
| Faecalibacterium | genus | 25.4 | 27.9 | 29 |
| [Bacteroides] coagulans | species | 41.4 | 44.7 | 41.1 |
| Hungatella | genus | 0.2 | 4 | 4 |
| Porphyromonadaceae | family | 20.4 | 16.3 | 20.8 |
| Adlercreutzia equolifaciens | species | 95 | 98 | 92.8 |
| Bifidobacterium catenulatum | species | 95.4 | 99.2 | 93.9 |
| Ruminococcaceae bacterium CC59_002D | strain | 88.9 | 94.4 | 94.4 |
| Aminicella | genus | 90.6 | 85.4 | 91.2 |
| Salinispirillum marinum | species | 34.2 | 30.9 | 37.5 |
| Eubacteriaceae | family | 61.8 | 66.2 | 58.8 |
| CCUG 54292 | species | 67.3 | 59.6 | 67.3 |
| Eubacterium ruminantium | species | 57.8 | 54.4 | 62.1 |
| Eubacterium | genus | 58.9 | 63.5 | 55.8 |
| Burkholderiales | order | 18.2 | 25.9 | 22.2 |
| Fibrobacteria | class | 18.3 | 26.1 | 22.5 |
| Bifidobacterium pseudocatenulatum | species | 87.2 | 93.8 | 86 |
| Coprococcus catus GD/7 | strain | 37.5 | 33.3 | 29.2 |
| Sutterellaceae | family | 19 | 16.3 | 24.7 |
| Clostridium sp. | species | 79.4 | 76.7 | 85.2 |
The amount of similarity caused me to want to cross check that this pattern is actually there and not the result of randomness.
Is this “seeing things” in the data or Seeing things?
I grab three other samples from different people who used CosmosId and plotted the difference between bacteria that were share in common. The number of bacteria in common with the family is 2.3x more than a random sample. Remember that many families will be found in almost all samples.
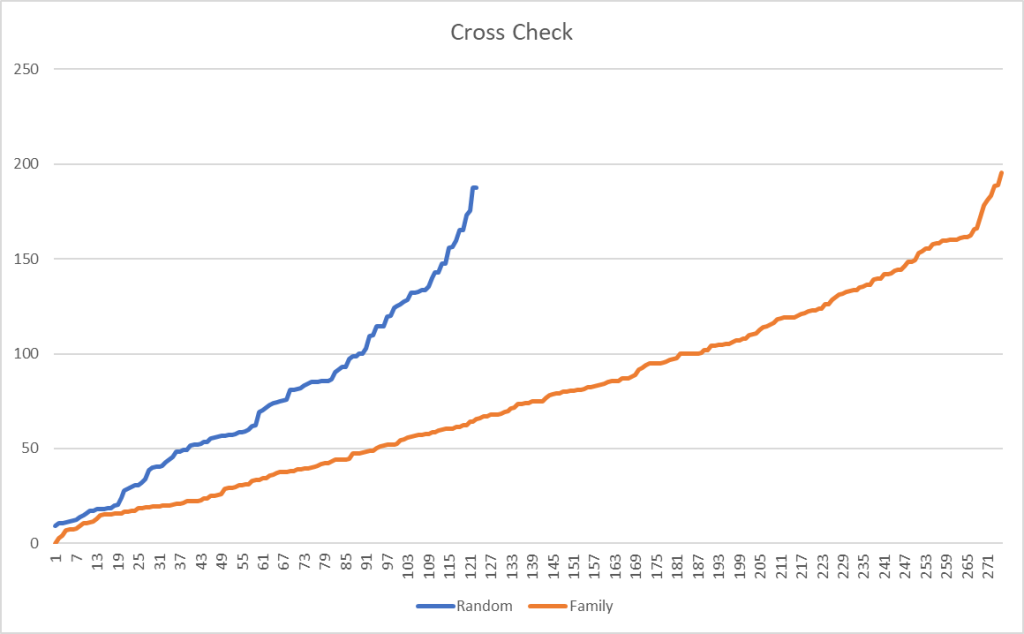
Filtering on species, results in a more dramatic change, with 2.68x more in common. Moving down to strains, it increased further to 3.64x more in common.
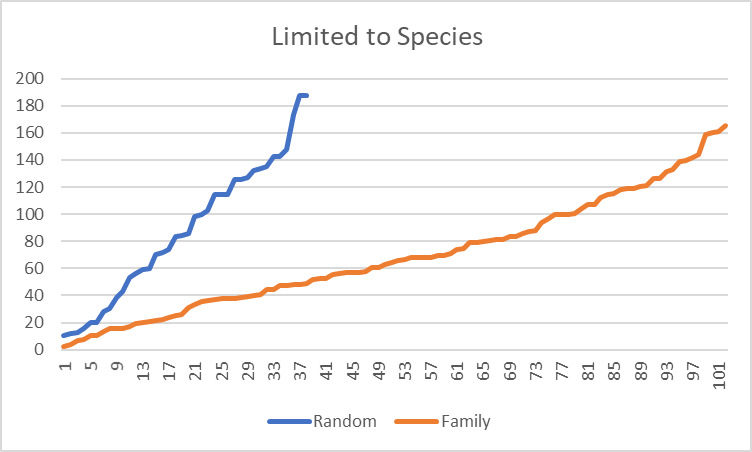
The shared diet, living space and DNA clearly results in a “family microbiome” pattern. It would be interesting to do an analysis of some couples that have lived together for at least 10 years (any volunteers?)
We will be using this shared pattern for a customized analysis. It may allow us to better isolated what may be contributing to the two issues cited above (and ignore items that may be high or low compared to the general population)
Identifying Probable Age related items
The microbiome changes with age, and for a number of bacteria we see what appear to be an age based signature.
Given that two are children and one is an adult, we can use how the microbiome changes with age to explain more of the results. The mother is either significantly higher to the levels seen by the kids, or significantly lower.
| Bacteria Name | Rank | Son | Mother | Daughter |
| Rhodocyclaceae | family | 0.3 | 10.8⬆️ | 3.5 |
| Lactobacillaceae | family | 6.4 | 12.5⬆️ | 2.4 |
| Fibrobacteres | phylum | 5.1 | 16.3⬆️ | 7.5 |
| Varibaculum cambriense | species | 3.5 | 16.6⬆️ | 8.8 |
| Clostridium clostridioforme | species | 0 | 11.3⬆️ | 3.8 |
| FCB group | clade | 8.7 | 19.2⬆️ | 4.7 |
| Bacteroidetes/Chlorobi group | clade | 8.2 | 19.1⬆️ | 4.2 |
| Bacteroidetes | phylum | 7.2 | 18.3⬆️ | 2.8 |
| Tyzzerella | genus | 0.7 | 17.3⬆️ | 3 |
| Fusobacterium | genus | 3 | 17.1⬆️ | 0.5 |
| Bacteroidetes | class | 7.3 | 20.2⬆️ | 3 |
| Bacteroidales | order | 7.3 | 20.2⬆️ | 3 |
| Faecalibacterium prausnitzii (Hauduroy et al. 1937) Duncan et al. 2002 | species | 22 | 40.1⬆️ | 21.1 |
| Corynebacteriaceae | family | 9.2 | 21.3⬆️ | 1.6 |
| Caseobacter | genus | 9 | 21.4⬆️ | 1.6 |
| Christensenella | genus | 69.4 | 90.3⬆️ | 68.2 |
| Christensenellaceae | family | 66.4 | 88.9⬆️ | 65.2 |
| Fibrobacteria | class | 18.3 | 26.1⬆️ | 22.5 |
| Bacteroides rodentium | species | 3.8 | 28.1⬆️ | 5.4 |
| Bryantella formatexigens | species | 5.4 | 28.8⬆️ | 0.7 |
| unclassified Erysipelotrichaceae (miscellaneous) | no rank | 28.8 | 5.8👇 | 34.6 |
| Lachnospira | genus | 3.2 | 30.3⬆️ | 1 |
| Parabacteroides distasonis CL09T03C24 | strain | 70 | 40👇 | 70 |
| Ruminococcus gnavus | species | 60.8 | 30.7👇 | 60.7 |
| Desulfovibrionaceae | family | 27.5 | 58.1⬆️ | 31.9 |
| Desulfovibrionales | order | 27.3 | 58⬆️ | 31.8 |
| Phocaeicola | genus | 57.6 | 25.6👇 | 55.3 |
| Eubacterium hallii | species | 58.4 | 90.5⬆️ | 69.8 |
| Butyrivibrio | genus | 9.2 | 36.9⬆️ | 4.2 |
| Intestinimonas | genus | 26.3 | 6.4👇 | 46.5 |
| Bilophila | genus | 31 | 71.⬆️1 | 37.5 |
The kids have two different issues:
- ASD/ADHD
- Constipation
If there was a third normal child in the same age range, picking bacteria would likely be trivial. What we have are just the difference. We do not know if having higher or lower is the cause of the issue. What we do have are bacteria to research and the mother.
| Bacteria Name | Rank | Son | Mother | Daughter |
| Actinobaculum | genus | 84.4 | 29.9 | 4.5👇 |
| Actinomadura | genus | 36.9 | 26.2 | 0.1👇 |
| Agathobaculum | genus | 0.7 | 1.4 | 21.7⬆️ |
| Alistipes | genus | 77.4 | 96.9 | 35.6👇 |
| Clostridioides | genus | 81.3⬆️ | 24.4 | 35.6 |
| Coprobacillus | genus | 81.3⬆️ | 4 | 0.7 |
| Discomyces | genus | 4.8👇 | 93.9 | 42.5 |
| Dorea | genus | 75.3 | 98.9 | 20.1👇 |
| Eggerthella | genus | 95.8⬆️ | 46 | 21.7 |
| Erysipelatoclostridium | genus | 0.8👇 | 9.4 | 6.1 |
| Escherichia | genus | 32.5👇 | 70.8 | 61.8 |
| Flavonifractor | genus | 0.1👇 | 15.4 | 20.4 |
| Holdemania | genus | 12.1👇 | 17.9 | 21.5 |
| Intestinibacter | genus | 26.8 | 6 | 58.3⬆️ |
| Lactobacillus | genus | 20.4 | 29.9 | 11👇 |
| Oscillibacter | genus | 44.7 | 43.5 | 80.3⬆️ |
| Pseudoflavonifractor | genus | 0.3👇 | 47.7 | 40.8 |
| Roseburia | genus | 4.7 | 28.7 | 57.7⬆️ |
| Ruminococcus | genus | 3.7👇 | 95.4 | 37.9 |
| Streptococcus | genus | 83.3⬆️ | 69.1 | 2.6 |
| Subdoligranulum | genus | 37.2 | 47.9 | 74.2⬆️ |
| Veillonella | genus | 0.4👇 | 14.9 | 9.4 |
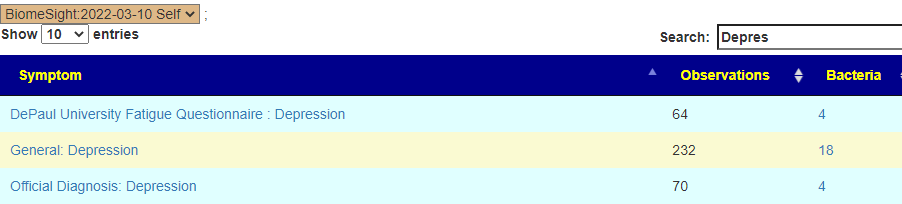
Looking at US Library of Medicine studies on ADHD, we have the following matches for the son:
Looking at Autism
- Alistipes (NCBI:239759 )
- Dorea (NCBI:189330 )
- Escherichia (NCBI:561 )
- Flavonifractor (NCBI:946234 )
- Lactobacillus (NCBI:1578 )
- Roseburia (NCBI:841 )
- Ruminococcus (NCBI:1263 )
- Veillonella (NCBI:29465 )
Nine shifts appear to match studies reported for the son.
Shifting to the daughter for constipation (using Functional constipation / chronic idiopathic constipation), we have just one
It is important to remember that the studies being referred to, were done on adults usually.
I was curious about Actinomadura because of the differences. It appears to occur in about 3% of samples, hence to get a reasonable sample to get relationships from, a clinical study would need to have 30/3% = 1000 participants and use a 16s processor that detects this bacteria (many do not). Worst yet, because of this rarity, we know of nothing that impacts it.
What to do when there is no information on changing a bacteria!!!
This actually opens the door to using bacteria-to-bacteria associations. We have 111 of them!
Looking at one with the highest positive associations, Desulfofarcimen, we see some items that will increase this bacteria (and by increasing that bacteria, Actinomadura should increase): cranberry polyphenols, Goji (berry,juice), walnuts, saccharin. Using all of the 111 interactions, we found only one thing that shows up as increasing Actinomadura (and many things to decrease), choline deficiency (i.e. reduce choline in diet). Items to avoid would be:
- jatropha curcas linn. (euphorbiaceae)
- nigella sativa seed (black cumin)
- oplopanax horridus(Devil’s Club)
- hypericin(St. John’s Wort)
- kefe cumin (laser trilobum l.)
- foeniculum vulgare (Fennel)
- lactobacillus bulgaricus (probiotics)
The only likely item would be yogurt containing lactobacillus bulgaricus).
Fortunately, most of the bacteria at the genus level has some studies on them.
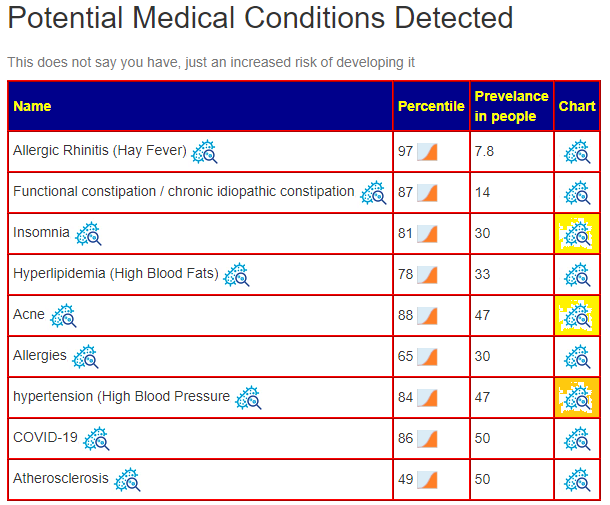
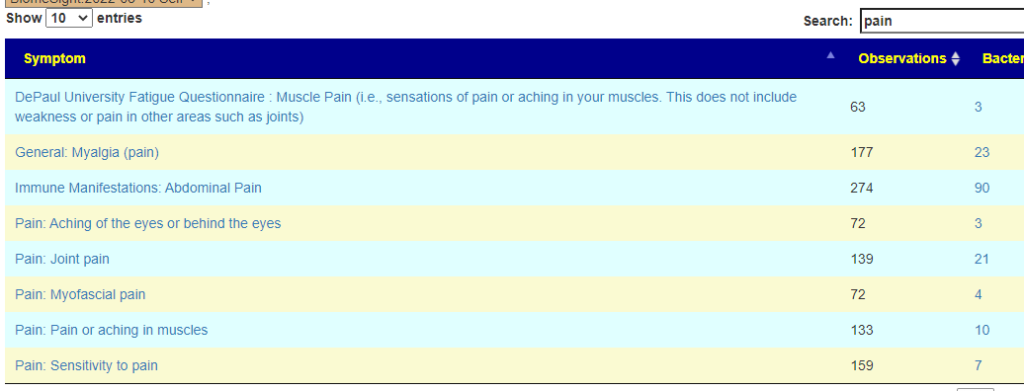
Symptom Prediction
This is from binary logistic regression which I have been working on but have not released. I picked the symptoms that appear viable for the reported issues. This is problematic because the regression was trained on adult data and not children — this is a mere exploration
The higher the number, the more likely, the lower the number the less likely.
| SYmptomName | Son | Mother | Daughter |
| Neurological: Short-term memory issues | -1.8 | -4.0 | -2.5 |
| Neurological: Disorientation | -5.1 | -7.2 | -6.6 |
| Immune Manifestations: Constipation | 0.5 | -0.2 | 1.2 |
| Neurological: High degree of Empathy before onset | -5.6 | -45.7 | -18.4 |
| Comorbid: Small intestinal bacterial overgrowth (SIBO) | -3.4 | 1.1 | -3.9 |
| Asymptomatic: Live in house with person with probable microbiome dysfunction | -7.2 | 1.5 | -11.2 |
| Neurocognitive: Difficulty paying attention for a long period of time | 0.7 | 0.3 | -0.3 |
| Neurocognitive: Can only focus on one thing at a time | 0.0 | -4.5 | -3.6 |
| Autism: Official Diagnosis | -1.8 | -3.8 | -2.5 |
| Comorbid: Constipation and Explosions (not diarrohea) | 0.9 | – ∞ | 1.7 |
| Comorbid: Constipation and Diarrohea (not explosions) | -2.4 | 0.3 | -0.5 |
The son is the most probable for:
- Autism: Official Diagnosis
- Neurological: Disorientation
- Neurocognitive: Difficulty paying attention for a long period of time (i.e. ADHD)
- Neurological: Short-term memory issues
- Neurocognitive: Can only focus on one thing at a time
The daughter is the most probable for:
- Immune Manifestations: Constipation
- Comorbid: Constipation and Explosions (not diarrohea)
The mother is most probable for:
- Asymptomatic: Live in house with person with probable microbiome dysfunction
- Comorbid: Small intestinal bacterial overgrowth (SIBO)
- Comorbid: Constipation and Diarrohea (not explosions)
While we had regression-training issues, using the “in the same microbiome family” comparison approach, the prediction largely modelled the actual symptoms reported.
Suggestions
For Daughter
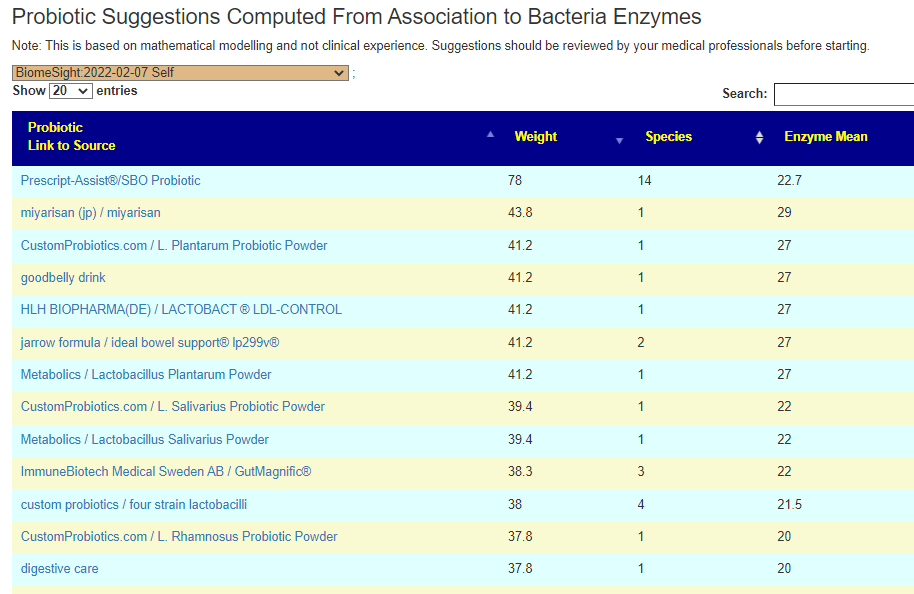
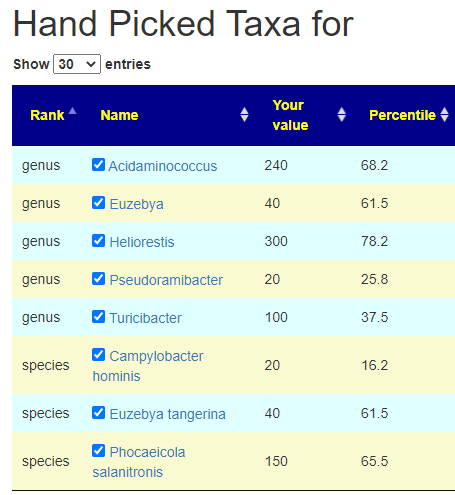
The daughter is the most challenging — only one match to PubMed studies that matches for constipation. I went to the sample, and made sure these two symptoms were marked. Then I went to the new feature:
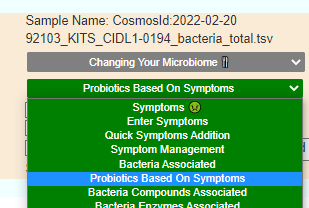
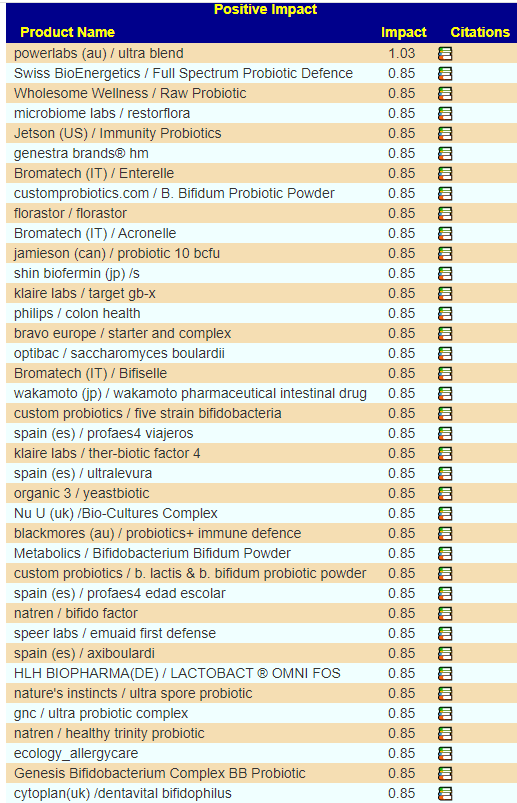
- #1 is o’donnell / flora-balance containing only bacillus laterosporus or brevibacillus laterosporus
- #2 is Prescript-Assist®/SBO Probiotic
- #3 is bacillus subtilis – which is available in many products
- Others high on the list contains bacillus subtilis
Next I flip over to data from Clinical Studies, The most studied was Lactobacillus rhamnosus GG (i.e. Culturelle®), followed by my old favorite Mutaflor (Escherichia coli strain Nissle 1917) which has limited availability Mutaflor (Canada, Australia, Finland, Germany) with Symbioflor-2 being a good alternative. Remember this is based on number of studies, not effectiveness.
Last, Ruminococcus is a bit of a challenge because both high and low values are associated in the literature — given the age, I deem it not safe to pursue changes in isolation.
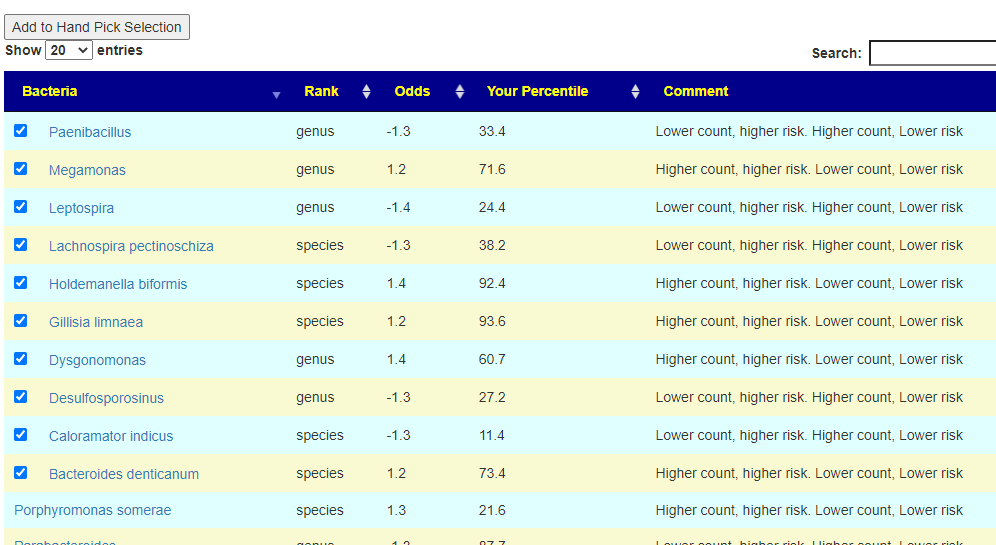
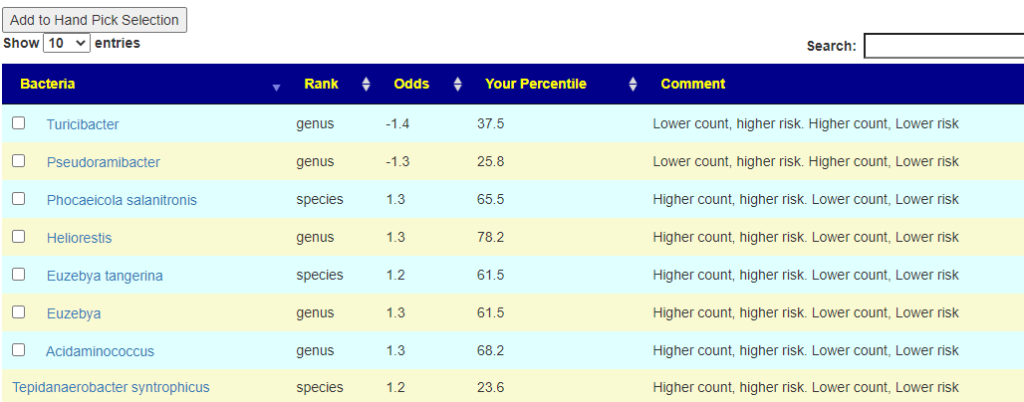
Looking at the heath analysis, everything looks good except for high Escherichia coli and Bacteroides fragilis. I suspect bad E.Coli is a factor for the constipation which suggests that the strong good E.Coli in Symbioflor-2 (or Mutaflor) would be beneficial. Using the various expert opinions (remember those are for adults!) we see 8,25, 76, 141 and 172 bacteria being selected.
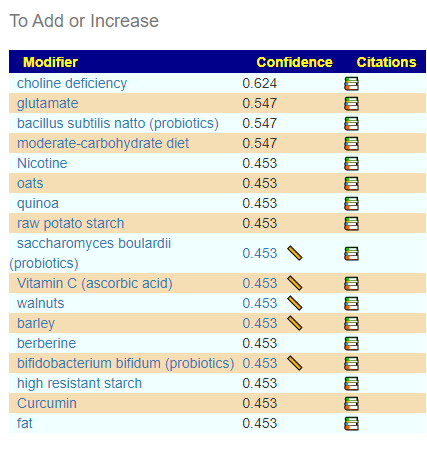
Going over to consensus, we see the following items outstanding (and likely path of least resistance to the daughter):
- pediococcus acidilactic (probiotic)
- bacillus coagulans (probiotics)
- bifidobacterium longum (probiotics)
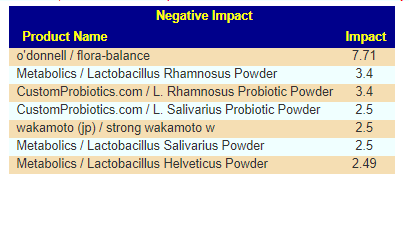
And the should-avoid including:
Bottom Line for Daughter
I would be inclined to bacillus subtilis alternating every 2 weeks with bacillus coagulans (probiotics) and bifidobacterium longum (probiotics) as the likely easiest to get. If not sufficient progress, then order Symboflor-2 from Germany (site that ships to the world)
For Son
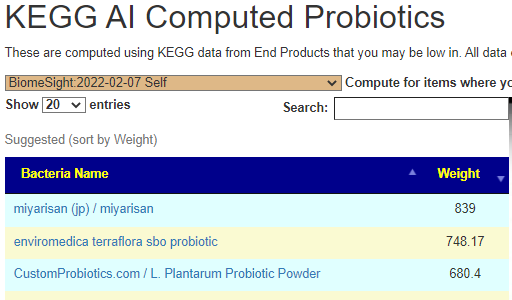
Doing the same pattern, adding symptoms and then doing KEGG computations for probiotics, we end up with a short list (in order): Prescript-Assist®/SBO Probiotic, enviromedica terraflora sbo probiotic, microbiome labs/ megasporebiotic, o’donnell / flora-balance — not a single lactobacillus or bifidobacterium in the list.
We have a reasonable list of bacteria identified above for Autism and ADHD, so I will run two advance suggestions with 15% selection and US Library of Medicine findings. The results were interesting – there was nothing in common with both sets of suggestions. The top items were:
- arabinoxylan oligosaccharides (prebiotic)
- soy
- inulin (prebiotic)
- lactobacillus plantarum (probiotics)
- lactobacillus rhamnosus gg (probiotics) ** this has been used in 3 studies with positive effects (Effect of probiotic supplementation on cognitive function in children and adolescents: a systematic review of randomised trials.)
- barley
Now over to the canned expert suggestions, Using the various expert opinions (remember those are for adults!) we see 7,10, 66, 121 and 225 bacteria being selected. With the following being good suggestions:
- pediococcus acidilactic (probiotic)
- inulin (prebiotic)
- lactobacillus plantarum (probiotics)
- lactobacillus reuteri (probiotics)
- soy
- Pulses
- high fiber diet
The avoid list top items were
This is very similar to the daughter and likely reflect the commonality of the microbiome.
The Mother
The meals for the kids are likely good suggestions for the entire family. There are not any active issues with the mother and the commonality of bacteria in all of the microbiome will lead to similar diet. I should mention that alcohol appears on the avoid list (not called out for the kids).
REMINDER
All of these are suggestions coming from mathematical models and not clinical experience. Suggestions should be reviewed by a knowledgeable medical professional before starting.
I am a computer scientist and a statistician. I am not licensed to practice medicine, and where I live has strict laws about ‘appearing to practice medicine’. What I can do for readers is to write a public blog (anonymous) from your data and back story as an education post on using the software and the statistics it produces. I cannot consult. The content should be reviewed by a medical professional before implementing.








Recent Comments