Tinnitus is not usually viewed as a microbiome issue. It was worth checking if it reaches our threshold for inclusion as defined in A new specialized selection of suggestions links. It did, hence this post
Study Populations:
| Symptom | Reference | Study |
| Neurological-Audio:Tinnitus (ringing in ear) | 1075 | 73 |
- Bacteria Detected with z-score > 2.6: found 129 items, highest value was 6.3
- Enzymes Detected with z-score > 2.6: found 493 items, highest value was 7.1
- Compound Detected with z-score > 2.6: found ZERO items
This is similar to Special Study: Histamine or Mast Cell Issues in finding no compounds, but the bacteria factor appears weaker and the enzymes is more.
Interesting Significant Bacteria
We have two dominant items that may be addressed by probiotics: Low Bifidobacterium and Low E.Coli
- E.Coli probiotics are Symbioflor-2 and Mutaflor.
- Bifidobacterium probiotics: we have 4 in the top group. Unfortunately none of these species are available at the retail level (that I am aware of). Checking interactions for these 4, there was no significant interactions found with common retail bifidobacterium species, just with the general genus.
Looking at Lactobacillus taiwanensis, there is a solid positive association with Bifidobacterium cuniculi and a weaker association with Bifidobacterium catenulatum subsp. kashiwanohense. There is also a weak association with two retail probiotic species: Bifidobacterium bifidum and Bifidobacterium animalis. It was interesting to note that there was no associations with any retail Lactobacillus species.
Bottom line appears to become: E.Coli probiotics, Bifidobacterium bifidum and Bifidobacterium animalis
All of the significant bacteria has too low levels.
| Bacteria | Reference Mean | Study | Z-Score |
| Bifidobacterium gallicum (species) | 3754 | 534 | 6.3 |
| Prevotella stercorea (species) | 6614 | 102 | 6 |
| Bifidobacterium subtile (species) | 82 | 32 | 5.3 |
| Escherichia coli (species) | 716 | 170 | 5.3 |
| Lactobacillus taiwanensis (species) | 85 | 12 | 5.3 |
| Catenibacterium mitsuokai (species) | 432 | 34 | 5.2 |
| Bifidobacterium cuniculi (species) | 81 | 28 | 5.1 |
| Enterobacteriaceae (family) | 8948 | 2622 | 5 |
| Bifidobacterium catenulatum subsp. kashiwanohense (subspecies) | 313 | 71 | 5 |
Interesting Enzymes
As above, all levels that were found significant had too little. I will leave it to the reader to go to Kyoto Encyclopedia of Genes and Genomes to learn about these enzymes (a steep learning curve).
| Enzyme | Reference Mean | Study Mean | Z-Score |
| [cysteine desulfurase]-S-sulfanyl-L-cysteine:[molybdopterin-synthase sulfur-carrier protein]-Gly-Gly sulfurtransferase (2.8.1.11) | 5326 | 2043 | 7.1 |
| gamma-L-glutamyl-L-cysteinyl-glycine:spermidine amidase (3.5.1.78) | 3212 | 1066 | 6.3 |
| gamma-L-glutamyl-L-cysteinyl-glycine:spermidine ligase (ADP-forming) [spermidine is numbered so that atom N-1 is in the amino group of the aminopropyl part of the molecule] (6.3.1.8) | 3212 | 1066 | 6.3 |
| tRNA-uridine13 uracil mutase (5.4.99.27) | 3671 | 1246 | 6.3 |
| donor:hydrogen-peroxide oxidoreductase (1.11.1.21) | 3052 | 869 | 6.2 |
| S-adenosyl-L-methionine:tRNA 5-(aminomethyl)-2-thiouridylate N-methyltransferase (2.1.1.61) | 3686 | 1305 | 6.1 |
| 5-oxo-L-proline amidohydrolase (ATP-hydrolysing) (3.5.2.9) | 10180 | 3505 | 6.1 |
| thioredoxin:protein disulfide oxidoreductase (dithiol-forming) (1.8.4.16) | 3812 | 1301 | 6.1 |
| acetyl-CoA:N6-hydroxy-L-lysine 6-acetyltransferase (2.3.1.102) | 719 | 143 | 6.1 |
| UDP-alpha-D-glucose:enterobactin 5′-C-beta-D-glucosyltransferase (configuration-inverting) (2.4.1.369) | 819 | 141 | 6.1 |
| S-adenosyl-L-methionine:tRNA (uracil54-C5)-methyltransferase (2.1.1.35) | 3817 | 1273 | 6.1 |
| L-methionine:2-oxo-acid aminotransferase (2.6.1.88) | 2771 | 815 | 6.1 |
Interesting Compounds
Nothing was found again!!!! In one sense this was a surprise, in another sense, it hints that the results found significant are not random.
Bottom Line
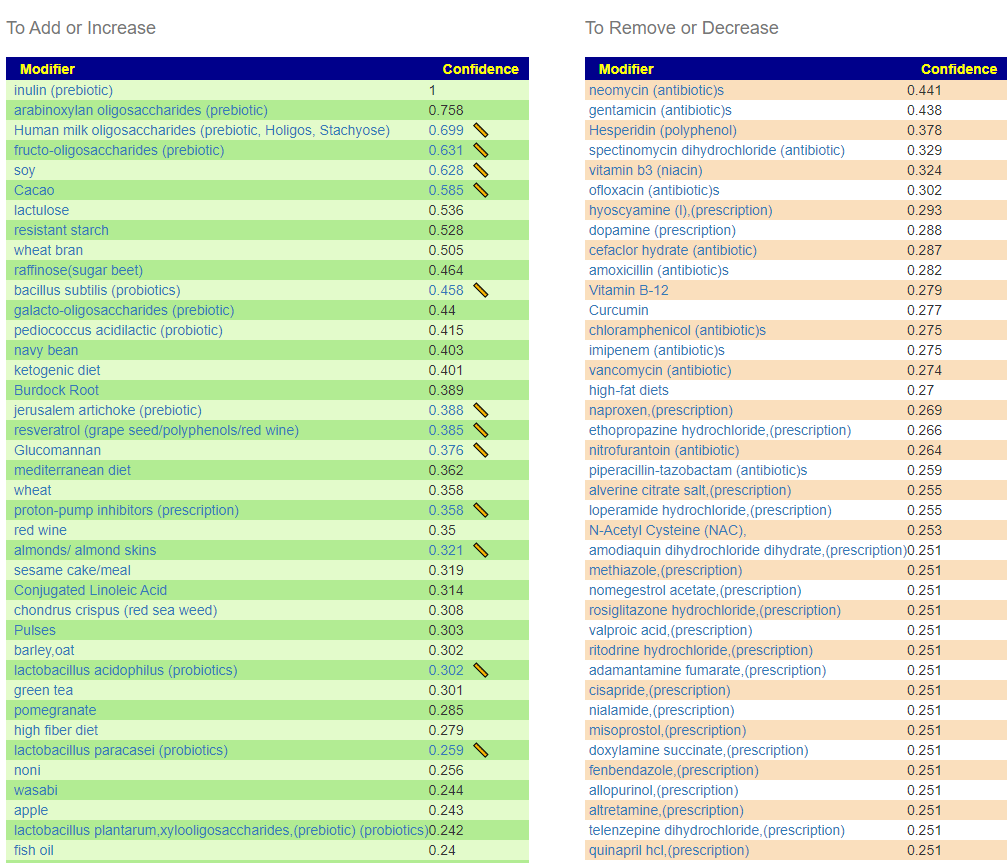
This was an interesting analysis because the dominant deficiencies were in genus that are available as probiotics. In reality, it points to just 3 probiotics: an E.Coli probiotic and two bifidobacterium. Food suggestions will be generated on Microbiome Prescription using an individual’s unique microbiome.
Social Feedback

I went and looked at these two in combinations and got a lot of bacteria in common
| Bacteria Name | Taxonomy rank |
| Nostoc | genus |
| Citrobacter | genus |
| Anaerococcus lactolyticus | species |
| Streptomycetales | order |
| Negativicoccus succinicivorans | species |
| Sutterella stercoricanis | species |
| Streptomycetaceae | family |
| Negativicoccus | genus |
| Lactococcus fujiensis | species |
| Actinomycetaceae | family |
| Prevotella disiens | species |
| Prosthecobacter | genus |
| Clostridium chartatabidum | species |
| Olivibacter | genus |
| Peptoniphilus asaccharolyticus | species |
| Butyricimonas synergistica | species |
| Campylobacterales | order |
| Epsilonproteobacteria | class |
| Campylobacteraceae | family |
| Pedobacter kwangyangensis | species |
| Staphylococcus pseudolugdunensis | species |
| Enterobacterales | order |
| Streptococcus milleri | species |
| Schaalia | genus |
| Enterobacteriaceae | family |
| Mitsuokella | genus |
| Gammaproteobacteria | class |
| Prevotellaceae | family |
| Clostridium cellulovorans | species |
| Schaalia naturae | species |
| Enterobacteriaceae incertae sedis | norank |
| Blochmannia | genus |
| Bulleidia | genus |
| Bifidobacterium cuniculi | species |
| Prevotella | genus |
| Escherichia | genus |
| Prevotella copri | species |
| Bifidobacterium gallicum | species |
| Escherichia coli | species |
| Prevotella stercorea | species |
| Prevotella paludivivens | species |
1 thought on “Special Studies: Tinnitus (ringing in ear)”
Comments are closed.