This person has been using microbiome prescription to reduce the symptoms with success and with objective measurements of improved microbiome. His MD is willing to prescribe antibiotics and the top three items (from hundreds possible) are all used by ME/CFS specialist — indicating that the model is in agreement with clinical experience of ME/CFS specialist (a.k.a. Cross-Validation).
This is a follow up to these prior posts:
- Dec 30, 2021. Rosacea, Circulation and mild CFS.
- Apr 27, 2022 Follow up Microbiome Analysis from a prior post
Why Follow Up Posts are important
The first item is simple, does the model and suggestion appear to work. Everything is theoretically computed. The second item is that encourages people to try suggestions
Foreword – and Reminder
I am not a licensed medical professional and there are strict laws where I live about “appearing to practice medicine”. I am safe when it is “academic models” and I keep to the language of science, especially statistics. I am not safe when the explanations have possible overtones of advising a patient instead of presenting data to be evaluated by a medical professional before implementing.
I cannot tell people what they should take or not take. I can inform people items that have better odds of improving their microbiome as a results on numeric calculations. I am a trained experienced statistician with appropriate degrees and professional memberships. All suggestions should be reviewed by your medical professional before starting.
Comparisons between Samples
First, I do not know the best way to compare samples — what I usually do is put all of the numbers side by side. Special attention needs to be paid to Lab Read Quality. A poorer read quality results in less bacteria being identified.
Lab Quality is a measure of the total number of bacteria counted. The processing of a sample may detect just 30,000 bacteria or 300,000 bacteria. This impacts the number of bacteria detected and also the accuracy of the measures.
| Criteria | 8/31/2021 | 12/3/2021 | 3/25/2022 | 8/11/2022 |
| Lab Read Quality | 7.8 | 3.6 | 6.2 | 5.5 |
| Bacteria Reported By Lab | 461 | 379 | 479 | 383 |
| Bacteria Over 99%ile | 7 | 5 | 3 | 3 |
| Bacteria Over 95%ile | 20 | 24 | 11 | 13 |
| Bacteria Over 90%ile | 32 | 40 | 21 | 23 |
| Bacteria Under 10%ile | 283 | 123 | 237 | 189 |
| Bacteria Under 5%ile | 222 | 66 | 143 | 107 |
| Bacteria Under 1%ile | 161 | 9 | 44 | 23 |
| Rarely Seen 1% | 3 | 2 | 14 | 7 |
| Rarely Seen 5% | 9 | 7 | 33 | 14 |
| Pathogens | 37 | 30 | 44 | 31 |
| Outside Range from JasonH | 4 | 4 | 7 | 7 |
| Outside Range from Medivere | 15 | 15 | 15 | 15 |
| Outside Range from Metagenomics | 6 | 6 | 8 | 8 |
| Outside Range from MyBioma | 7 | 7 | 7 | 7 |
| Outside Range from Nirvana/CosmosId | 18 | 18 | 23 | 23 |
| Outside Range from XenoGene | 5 | 5 | 7 | 7 |
| Outside Lab Range (+/- 1.96SD) | 14 | 9 | 6 | 8 |
| Outside Box-Plot-Whiskers | 41 | 58 | 38 | 33 |
| Outside Kaltoft-Møldrup | 211 | 100 | 123 | 111 |
| Condition Est. Over 99%ile | 0 | 0 | 0 | 0 |
| Condition Est. Over 95%ile | 4 | 3 | 1 | 1 |
| Condition Est. Over 90%ile | 9 | 6 | 5 | 7 |
| Enzymes Over 99%ile | 17 | 19 | 30 | 10 |
| Enzymes Over 95%ile | 105 | 82 | 219 | 68 |
| Enzymes Over 90%ile | 139 | 126 | 296 | 183 |
| Enzymes Under 10%ile | 783 | 369 | 514 | 645 |
| Enzymes Under 5%ile | 542 | 186 | 264 | 423 |
| Enzymes Under 1%ile | 271 | 37 | 49 | 86 |
| Compounds Over 99%ile | 33 | 28 | 62 | 47 |
| Compounds Over 95%ile | 140 | 127 | 231 | 254 |
| Compounds Over 90%ile | 346 | 307 | 298 | 338 |
| Compounds Under 10%ile | 310 | 227 | 297 | 308 |
| Compounds Under 5%ile | 211 | 111 | 224 | 173 |
| Compounds Under 1%ile | 132 | 47 | 67 | 65 |
The next table is also very dependent of Lab Read Quality. The apparent improvement on 12/3/2021 is likely artificial because the counts are low due to low read quality.
| 8/31/2021 | 12/3/2021 | 3/25/2022 | 8/11/2022 | |
| Percentile | Genus | Genus | Genus | Genus |
| 0 – 9 | 73 | 24 | 51 | 51 |
| 10-19 | 15 | 18 | 32 | 24 |
| 20 – 29 | 12 | 13 | 18 | 12 |
| 30 – 39 | 4 | 10 | 9 | 14 |
| 40 – 49 | 6 | 8 | 9 | 3 |
| 50 – 59 | 4 | 8 | 7 | 2 |
| 60 – 69 | 4 | 4 | 9 | 3 |
| 70 – 79 | 7 | 10 | 7 | 10 |
| 80 – 89 | 7 | 4 | 8 | 5 |
| 90 – 99 | 14 | 18 | 8 | 8 |
| 8/31/2021 | 12/3/2021 | 3/25/2022 | 8/11/2022 | |
| Percentile | Species | Species | Species | Species |
| 0 – 9 | 87 | 29 | 57 | 58 |
| 10-19 | 24 | 21 | 29 | 24 |
| 20 – 29 | 14 | 15 | 21 | 16 |
| 30 – 39 | 10 | 16 | 14 | 14 |
| 40 – 49 | 2 | 6 | 14 | 3 |
| 50 – 59 | 12 | 9 | 17 | 10 |
| 60 – 69 | 9 | 10 | 10 | 7 |
| 70 – 79 | 8 | 9 | 14 | 7 |
| 80 – 89 | 7 | 15 | 5 | 4 |
| 90 – 99 | 11 | 13 | 10 | 9 |
So how to interpret this wall of numbers? People can cherry-pick the numbers to say improvement or no improvement. The difference of lab read quality is a big factor because they impact the count for most of the items above. The Outside Box-Plot-Whiskers numbers show continued improvement. In short, the changes shown were less than I was hoping to see.
There is one more method of comparison — using special studies. In this case we see the average matches. Doing a little math, the expected drop of percentage due to lab quality size between 8/31/2021 and 8/11/2022 is a 10% drop. Those that exceeded 20% are color with 😊 below. Nothing became 10% worse. Note that the 😊 also agrees with comparing to 3/25/2022 (the prior sample). Other items remained unchanged. Items reported by this person are 😧 – Strong issue, 😟 – a bit of an issue
| Study | 8/31/2021 | 12/3/2021 | 3/25/2022 | 8/11/2022 | Average |
| Inflammatory bowel disease | 49 | 32 | 47 | 47 | 43.8 |
| Small intestinal bacterial overgrowth (SIBO) 😟 | 51 | 29 | 48 | 34😊 | 40.5 |
| Allergic Rhinitis (Hay Fever) 😧 | 41 | 27 | 45 | 41 | 38.5 |
| Autism | 45 | 34 | 44 | 27😊 | 37.5 |
| COVID19 (Long Hauler) | 40 | 24 | 43 | 29😊 | 34.0 |
| Irritable bowel syndrome 😟 | 40 | 22 | 44 | 30😊 | 34.0 |
| Alcohol intolerance or Medication sensitivities | 43 | 23 | 39 | 29😊 | 33.5 |
| Histamine or Mast Cell issues 😧 | 44 | 17 | 43 | 24😊 | 32.0 |
| Post-exertional malaise 😧 | 36 | 23 | 40 | 29😊 | 32.0 |
| ME/CFS without IBS 😟 | 39 | 22 | 36 | 30😊 | 31.8 |
| Poor gut motility | 42 | 21 | 40 | 24😊 | 31.8 |
| Brain Fog 😧 | 37 | 23 | 33 | 33 | 31.5 |
| Depression | 32 | 27 | 32 | 34 | 31.3 |
| Allergies And Food Sensitivity 😧 | 38 | 23 | 36 | 25😊 | 30.5 |
| Cold Extremities 😧 | 42 | 23 | 33 | 22😊 | 30.0 |
| Intolerance of Extremes of Heat and Cold 😟 | 38 | 16 | 39 | 27😊 | 30.0 |
| ME/CFS with IBS 😟 | 36 | 21 | 36 | 27 | 30.0 |
| Bloating 😧 | 37 | 22 | 34 | 26😊 | 29.8 |
| General Fatigue 😧 | 34 | 20 | 31 | 34 | 29.8 |
| Unrefreshed sleep | 31 | 22 | 36 | 28 | 29.3 |
| Constipation | 37 | 21 | 29 | 25😊 | 28.0 |
| High Anxiety 😟 | 33 | 17 | 33 | 29 | 28.0 |
| Easily irritated 😟 | 32 | 16 | 34 | 23😊 | 26.3 |
| Tinnitus (ringing in ear) 😟 | 24 | 15 | 29 | 22 | 22.5 |
| Chronic Fatigue Syndrome (CFS/ME) 😧 | 25 | 21 | 20 | 23 | 22.3 |
| Average | 37.8 | 22.4 | 37.0 | 28.9 | 31.5 |
| Lab Quality | 7.8 | 3.6 | 6.2 | 5.5 |
What is my conclusion? Most of the measures above deteriorates into noise with the exception of data from Special Studies, where we seen improvement in many measures, but not all. In one real statistical sense this makes sense: many are based on common sense and the ones showing clear improvement on statistical significance.
Going Forward
For most of my prior posts used the logical reasoning and clinical studies (which used different labs and software than the samples that I was looking at). With the special studies, we have upped our game (potentially) – the bacteria deemed significant were determined by the same lab and software of our sample, plus the study sizes was much larger than published clinical studies — hence better detection.
To build the consensus I will use the special studies, I filtered to reported issues and high percentage of matches, namely:
- Allergic Rhinitis (Hay Fever)
- Histamine or Mast Cell issues
- Post-exertional malaise
- Brain Fog
- Allergies And Food Sensitivity
- Cold Extremities
- General Fatigue
Remember that most of the special studies found that infrequent bacteria with a low value was what was statistically significant. This is turning the usual logic on it’s head. As I state, this is all experimental but based on studies and statistics.
The top suggestions are below
- navy bean
- Pulses ( dry peas, beans, lentils, chickpeas, etc)
- arabinoxylan oligosaccharides (prebiotic)
- non-starch polysaccharides
- red wine
- wheat bran
- proton-pump inhibitors (prescription)
- l-citrulline
- xylan (prebiotic)
- pea (fiber, protein)
Antibiotics
As expected, most antibiotics and prescription drugs are to be avoided. A few with positive impact includes:
- macrolide ((antibiotic)s) – used by Dr. Cecile Jadin in treating ME/CFS
- sulfonamide (antibiotic)s – suggested for treated ME/CFS [2019]
In terms of generic suggestions, rifaximin (antibiotic)s is by far the top antibiotics, cited here on Health Rising: Rifaxamin – citing use by Dr. Teitelbaum, Dr. Peterson, De De Meirleir and Dr. Myhill (all ME/CFS specialists).
In short, all of the top suggested antibiotics are applicable. My personal approach would be do all three of them in a pulse manner a la Jadin, 10 days on, 20 days off and then move to the next one.
Both above and generic suggestions have proton-pump inhibitors (prescription) being the top choice for other prescription drugs.
Probiotics
The top probiotics list have the usual dilemma: both e.coli probiotics and lactobacillus probiotics. It’s a dilemma because they tend to be hostile to each other. My typical rotation resolution would be 2 weeks of each and then move to the next:
- saccharomyces boulardii (probiotics)
- e.coli probiotics (take it with d-ribose, on the to take list and a food for e.coli)
- bifidobacterium
- lactobacillus
- Equilibrium and/or Prescript Assist
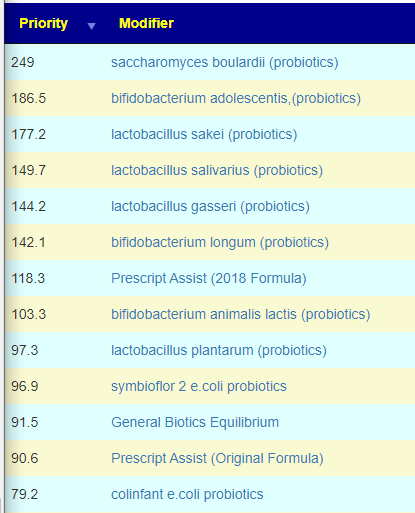
I should note that some are strong to be avoided (watch out for mixtures!!!)
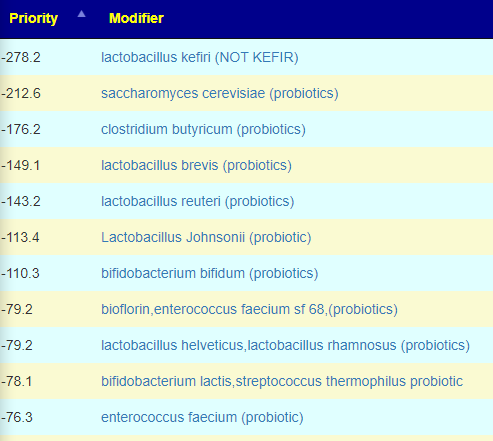
KEGG Suggestions
The KEGG suggestions top items were the bacteria found in Equilibrium and Prescription Assist, except for the top choice, Escherichia coli. A probiotic suggested by Dr. Myhill, a ME/CFS specialist in the UK. The next common conventional items are
Bottom Line
The suggestions above were done solely from special studies. The key question is are they reasonable? I would say yes based on the antibiotics suggestions — all of them have been reported to help ME/CFS patients. We also have agreement between KEGG probiotics and these suggestions.
There is a potential conceptual symmetry between the two approaches (working off extremes and using special studies that are often dealing with rare low bacteria). Bacteria influences each other in very complex ways.
The full list of suggestions is available above.
1 thought on “ME/CFS Follow Up Microbiome Samples”
Comments are closed.