This is a look at Small intestinal bacterial overgrowth (SIBO) from the aspect of the microbiome. Whatever is happening in the small intestine flows out of it and impacts downstream systems. Items that modifies any microbiome abnormalities will likely impact the small intestine.
This post is a parallel to my earlier post Multiple Chemical Sensitivity (MCS) – A Cause Found?. We will not assume anything. Just let the data speak.
We have 181 samples annotated with SIBO. Ombre Labs: 57, uBiome: 57 and Biomesight: 57.
Literature Review
I found one study that does a very good job
Association between Gut Dysbiosis and the Occurrence of SIBO, LIBO, SIFO and IMO [2023] states “Bacteria characteristic of small intestinal bacterial overgrowth include Streptococcus, Staphylococcus, Bacteroides, and Lactobacillus. “
Microbiome Associations
We have two bacteria that are reported significant in at least 2 of the 3 labs:
- Bacteroides caccae (species)
- Holdemania (genus)
A total of 31 bacteria (P < 0.001) was flagged as being significant and added to our collection.
We then look at Percentile and found only Bifidobacterium adolescentis showing a significant shift.
The complete list is below. Note that almost everything is High with SIBO.
| ShiftIs | tax_name | tax_rank |
| High | Flavobacteriia | class |
| High | Bacteroidia | class |
| High | Spongiibacteraceae | family |
| High | Lachnospiraceae | family |
| High | Bacteroidaceae | family |
| High | Flavobacteriaceae | family |
| High | Rhodocyclaceae | family |
| High | Anaerolinea | genus |
| High | Ethanoligenens | genus |
| High | Trabulsiella | genus |
| High | Bacteroides | genus |
| High | Hyphomicrobium | genus |
| High | Myroides | genus |
| high | Holdemania | genus |
| High | Pseudoflavonifractor | genus |
| High | Adlercreutzia | genus |
| High | Bacteroidales | order |
| High | Flavobacteriales | order |
| High | Oceanospirillales | order |
| High | Nitrosomonadales | order |
| High | Bifidobacterium thermophilum | species |
| High | Hyphomicrobium aestuarii | species |
| High | Anaerolinea thermolimosa | species |
| High | Thermodesulfovibrio thiophilus | species |
| High | Bacteroides sp. dnLKV9 | species |
| High | Bacteroides sp. 35AE37 | species |
| High | Bacteroides caccae | species |
| High | Adlercreutzia equolifaciens | species |
| High | Blautia obeum | species |
| High | Pseudoflavonifractor capillosus | species |
| Low | Bifidobacterium adolescentis | species |
We have only one clear agreement with the literature: Bacteroides with a large number of species deemed significant.
Compounds Produced
SIBO is usually detected by breath tests. The question is whether the chemicals detected in the breath is associated with the compounds that bacteria produced. What we found is below. The top 4 product in this list are the likely suspect for the first stage. These likely contributed to the chemicals detected. In particular, N-Acyl-L-homoserine abundance is associated with “Over 50 Gram-negative bacteria species (including several pathogenic species) use AHLs as autoinducers and the means of their communication in Quorum sensing” In other words it is not the source, but sends signals to the source bacteria to release the chemicals detected on the breath.
Trying to “find” the specific bacteria is not practical
- 📚4543 different bacteria produces Hydrogen (just click to see the list)
- 📚3817 different bacteria produces Hydrogen Sulfide
- 📚622 different bacteria produces Methane.
| CompoundName | percentile | Obs |
| Methylaminoacrylate C20255 – has NH2 and Hydrogen | 61.5 | 129 |
| Aminoacrylate C20253 – has NH2 and Hydrogen | 61.5 | 129 |
| D-Erythritol 1-phosphate C21294 | 60.7 | 134 |
| N-Acyl-L-homoserine C18061 – a signaling molecule | 60.3 | 126 |
| 2-O-[6-O-Octanoyl-alpha-D-glucopyranosyl-(1->6)-alpha-D-glucopyranosyl]-D-glycerate | 39.8 | 44 |
| (2R,3R)-Dihydroflavonol | 38.6 | 67 |
| 6-Phospho-2-dehydro-D-gluconate | 36.9 | 41 |
| p-Hydroxybenzyl alcohol | 34.8 | 56 |
| 4-Hydroxybenzaldehyde | 34.8 | 56 |
Bottom Line
I have made the bacteria found above available on the site under Research tab

The page resulting looks like this:
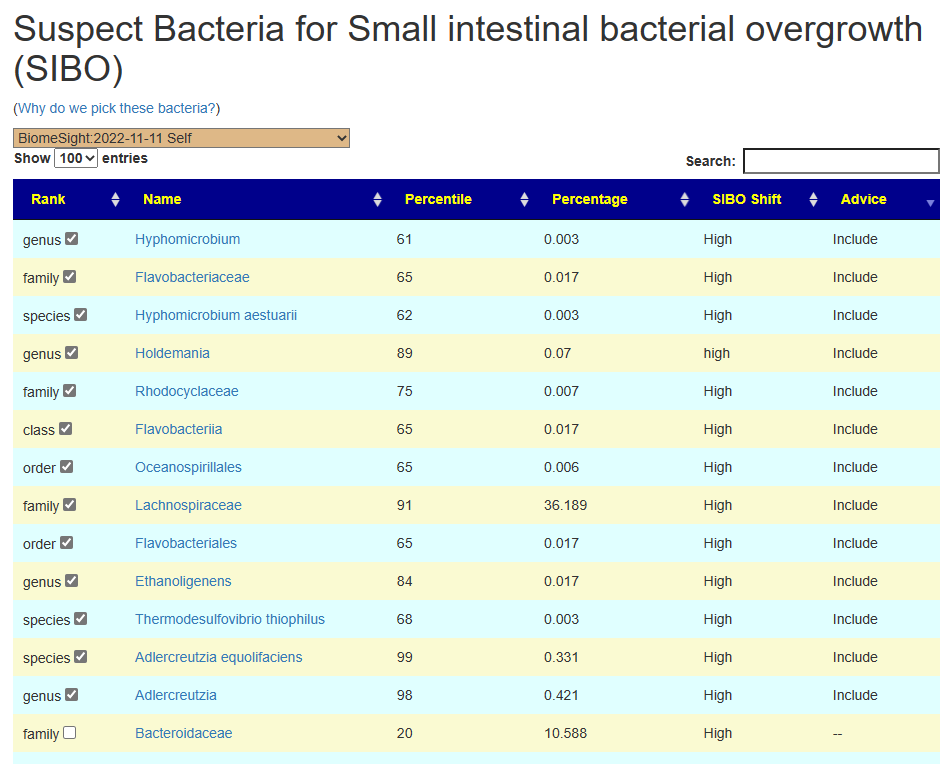
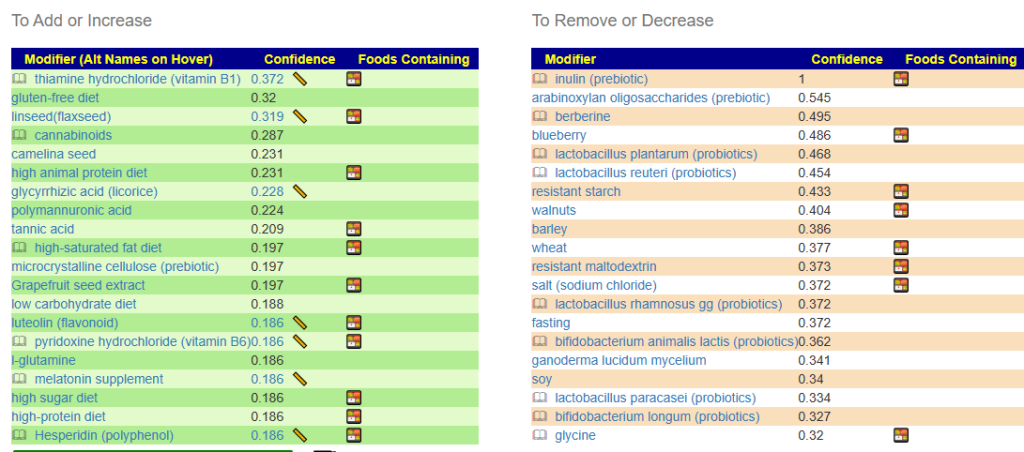
With results like shown below. Note that on the Add with have gluten-free and on the Remove: wheat, barley.
Recent Comments