This sketch is suggested for all studies looking for associations of bacteria to any thing else. I will illustrate a sample results based on a sample of 100 people with self-declared Autism against a reference population at the end. The need for a better standard came out of this post, Microbiome Statistics for Dummies. As a FYI, applying all of the methods below had over 200 finding with P < 0.01 from the test sample.
Caveat emptor: This is based on 16s samples using self-declaration for Autism. The purpose of this post is not to report on microbiome shifts for Autism, but to show different statistics that should be used, but tend to be ignored.
Using Odds Ratios
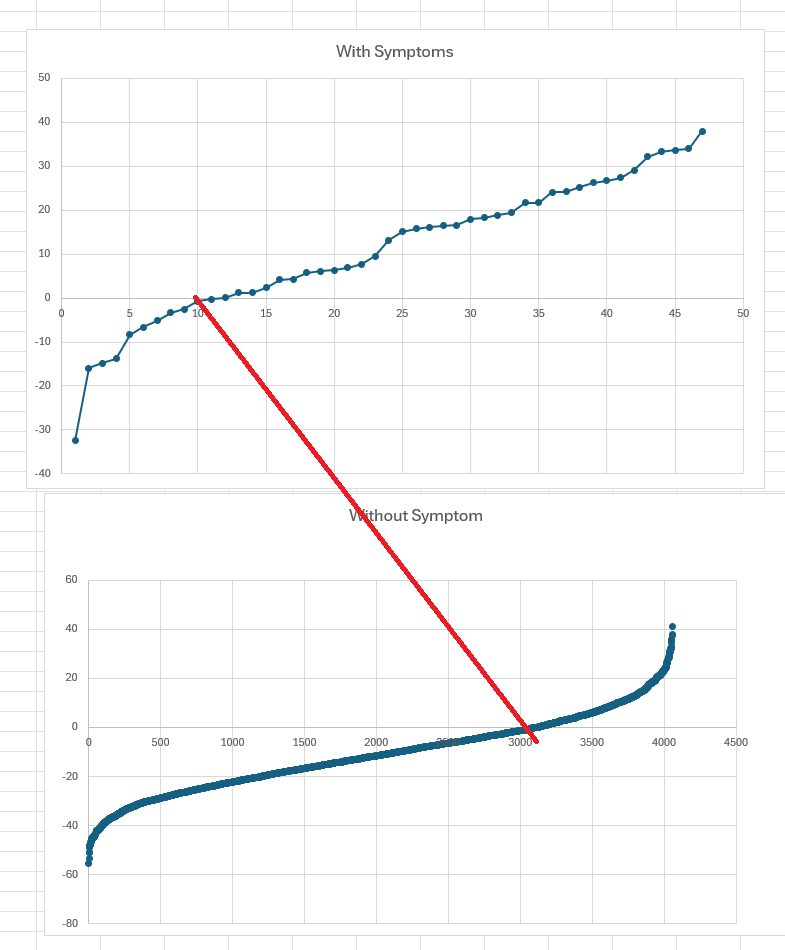
The charts below used naïve odds ratio computation (ignoring Probability) of people with declared symptom and those without this symptoms. If you use Zero(0) as a threshold, we correctly predicted 74% of the time for those with symptoms and those without symptoms.
Revised Reports
After checking (and repairing) some calculations, I am planning to publish every few days another snapshot such as the ones below. I will update this page with each publication.
- Odds Ratio Snapshot: Attention deficit hyperactivity disorder (ADHD)
- Odds Ration Snapshot: Bloating
- Odds Ratio Snapshot: Depression – With literature reference showing success with the top common probiotic.
- Odds Ratio Snapshot: Gluten Sensitivity (Non-Celiac)
- Odds Ratio Snapshots: Histamine or Mast Cell issues
- Odds Ratio Snapshot: Long COVID
- Odds Ratio Snapshot: ME/CFS without IBS
- Odds Ratio Snapshot: ME/CFS with IBS
- Odds Ratio Snapshots: Neurocognitive: Brain Fog
- Odds Ratios for Neurological-Audio: hypersensitivity to noise
- Odds Ratio Snapshots: Official Diagnosis: Chronic Fatigue Syndrome (CFS/ME)
- Odds Ratio Snapshot: Official Diagnosis: Mast Cell Dysfunction
- Odds Ratio Snapshot: Photophobia (Light Sensitivity)
- Odds Ratio Snapshot: Restless Legs
- Odds Ratio Snapshot: Small intestinal bacterial overgrowth (SIBO)
- Odds Ratio Snapshot: Tinnitus (ringing in ear)
- Odds Ratio Snapshots: Unrefreshed sleep
- Odds Ratio Snapshot: Gluten Sensitivity (Non-Celiac)
Criteria
Measures must have the ability to compute statistical significance. Measures should be multi-faceted and not mono-faceted (i.e. comparing average only). The following measures are proposed as a new standard:
- Odds Ratio based on the Median of the targeted population.
- Odds Ratio based on the Incidence of being seen between populations
- Odds Ratio based on the Median of the reference population.
- Difference of Median between target and general population
- Difference of Means between target and general population
- Difference of Skews between target and general population
Odds Ratio
To compute an odds ratio (OR) and assess its statistical significance, you
- (1) calculate the OR from a 2×2 table, then
- (2) compute a confidence interval (CI) and/or a p‑value
- via chi‑square test, or Fisher’s exact test
So we have for the three odds ratio cited above:
- Median Odds Ratio When Detected
- Numbers above/below the Median of the target/reference populations
- For Reference Population
- For Target Population
- Numbers above/below the Median of the target/reference populations
- Incidence Odds Ratio When Detected
- Number where it was detected, Number where it was not detected
- For Reference Population
- For Target Population
- Number where it was detected, Number where it was not detected
Difference of Skews
While this is not normally done, given the very high skewness (20+) seen with bacteria, it is a worthwhile investigation when sample sizes are sufficient. A description of how to do this is below.
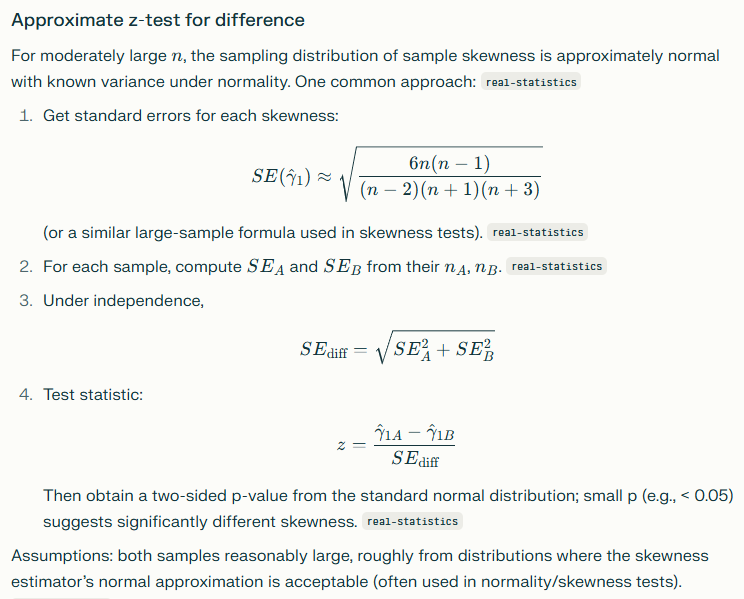

Difference of Means
Given the high skewness of most bacteria, this is a well known standard approach will typically underestimate the significance.
Real Data using 100 Autism People
Bacteria Incidence P < 0.01
This first table caused me to do a double take because Bifidobacterium is generally low in Autism and yet we have a table with the odds of having Autism being increased with certain Bifidobacterium. The key factor to remember is that Bifidobacterium is rarely reported with Autism — but when it is reported, the amounts tend to be higher; additionally, certain species tend to be a lot more common with Autism.
To verify my computations, I also included the percentage of time it is seen with Autism and without. These Bifidobacterium are far more common to appear in samples.
| tax_name | rank | Incidence Odds RATIO | Chi2 | Symptom % Seen | Reference % Seen |
| Bifidobacterium catenulatum subsp. kashiwanohense | subspecies | 2.8 | 39.80888 | 54 | 19 |
| Bifidobacterium angulatum | species | 2.8 | 32.81414 | 45 | 16 |
| Hungateiclostridium | genus | 2.4 | 17.91463 | 30 | 12 |
| Hungateiclostridiaceae | family | 2.4 | 17.7539 | 30 | 12 |
| Bifidobacterium scardovii | species | 2.1 | 13.64735 | 34 | 16 |
| Clostridium chartatabidum | species | 1.9 | 14.42216 | 48 | 24 |
| Bifidobacterium catenulatum PV20-2 | strain | 1.9 | 15.85311 | 63 | 33 |
| Bifidobacterium catenulatum | species | 1.9 | 15.01648 | 64 | 35 |
| Parascardovia | genus | 1.8 | 9.703312 | 35 | 19 |
| Bifidobacterium cuniculi | species | 1.8 | 9.141149 | 34 | 18 |
| Bifidobacterium gallicum | species | 1.7 | 12.55468 | 82 | 48 |
| ant endosymbionts | clade | 1.7 | 7.530999 | 39 | 23 |
| Candidatus Blochmanniella | genus | 1.7 | 7.530999 | 39 | 23 |
| Enterobacteriaceae incertae sedis | no rank | 1.6 | 6.782294 | 39 | 24 |
| Enterobacter hormaechei | species | 1.6 | 6.962239 | 41 | 25 |
| Moorella group | norank | 1.6 | 8.616342 | 66 | 42 |
| unclassified Bacteroidetes Order II. | order | 1.6 | 8.740627 | 75 | 48 |
| Bifidobacterium indicum | species | 1.6 | 8.410521 | 76 | 49 |
Using Symptom Median P < 0.01
We have 152 bacteria identified. Bifidobacterium species again featured. For brevity, I show only the top 25 below. Remember using the Medium of those with Autism, 50% of Autistic people will be below, and 50% above — there is no need to show those counts.
Note that the top line is saying if the amount of Bifidobacterium is above 0.85 that the odds ratio is just .26 (greatly reduced). The Odds Ratio applies to those with the reported amount above the Median.
REMINDER: We are looking at only samples finding bacteria.
| tax_name | Rank | Median | Odds Ratio | Chi2 | Below | Above |
| Bifidobacterium | genus | 0.8505 | 0.26 | 49.3 | 3114 | 817 |
| Actinomycetota | phylum | 1.0869 | 0.28 | 44.5 | 3126 | 872 |
| Bifidobacteriaceae | family | 0.764 | 0.28 | 43.4 | 3075 | 870 |
| Bifidobacteriales | order | 0.764 | 0.28 | 43.4 | 3075 | 870 |
| Bifidobacterium catenulatum | species | 0.01 | 0.29 | 39 | 1148 | 331 |
| Bifidobacterium asteroides | species | 0.005 | 0.29 | 36.8 | 872 | 255 |
| Bifidobacterium subtile | species | 0.0065 | 0.33 | 31.1 | 1093 | 357 |
| Actinomycetes | class | 0.71595 | 0.34 | 30.7 | 2978 | 1013 |
| Bifidobacterium cuniculi | species | 0.006 | 0.31 | 30.7 | 598 | 188 |
| Parascardovia | genus | 0.004 | 0.32 | 30.6 | 608 | 192 |
| Leyella | genus | 0.005 | 0.35 | 28.4 | 1314 | 454 |
| Leyella stercorea | species | 0.005 | 0.35 | 28.4 | 1314 | 454 |
| Bacteroides rodentium | species | 0.0695 | 2.72 | 26.1 | 1060 | 2878 |
| Bifidobacterium gallicum | species | 0.084 | 0.37 | 24.9 | 1508 | 559 |
| Moraxellales | order | 0.004 | 0.37 | 24.7 | 1597 | 595 |
| Moraxellaceae | family | 0.004 | 0.37 | 24.7 | 1597 | 595 |
| Eukaryota | superkingdom | 0.004 | 0.37 | 24.6 | 1089 | 401 |
| Phocaeicola paurosaccharolyticus | species | 0.025 | 2.62 | 24.1 | 1080 | 2828 |
| Geopsychrobacteraceae | family | 0.017 | 0.39 | 22.7 | 1400 | 540 |
| Desulfuromusa | genus | 0.017 | 0.39 | 22.7 | 1400 | 540 |
| Burkholderiales genera incertae sedis | no rank | 0.018 | 0.38 | 22.4 | 1024 | 393 |
| Desulfuromonadales | order | 0.015 | 0.39 | 22.4 | 1575 | 614 |
| Desulfuromonadia | class | 0.015 | 0.39 | 22.3 | 1574 | 614 |
| Desulfuromonadaceae | family | 0.016 | 0.39 | 21.9 | 1446 | 568 |
| Psychrobacter | genus | 0.003 | 0.39 | 21.5 | 1250 | 493 |
Using Reference Median P < 0.01
In this case, we have 89 bacteria. Bifidobacterium is very common again. To clarify matters a little: If the sample has Bifidobacterium reported and the amount is over 0.115, the odds of this person having Autism is 3.66.
The Odds Ratio applies to those with the reported amount above the Median
| tax_name | Rank | Reference Median | Odds Ratio | Chi2 | Symptom Below | Symptom Above |
| Bifidobacteriales | order | 0.116 | 3.71 | 32.1 | 21 | 78 |
| Bifidobacteriaceae | family | 0.116 | 3.71 | 32.1 | 21 | 78 |
| Bifidobacterium | genus | 0.115 | 3.66 | 31.3 | 21 | 77 |
| Actinomycetes | class | 0.175 | 3.35 | 28.5 | 23 | 77 |
| Caloramator indicus | species | 0.005 | 0.09 | 25.8 | 34 | 3 |
| Bifidobacterium gallicum | species | 0.009 | 3.37 | 23.9 | 19 | 64 |
| Actinomycetota | phylum | 0.23 | 2.85 | 22.5 | 26 | 74 |
| Hathewaya histolytica | species | 0.159 | 0.39 | 18.3 | 71 | 28 |
| Hathewaya | genus | 0.159 | 0.39 | 18.3 | 71 | 28 |
| Anaerovibrio lipolyticus | species | 0.028 | 0.38 | 17.1 | 63 | 24 |
| Rhodothermota | phylum | 0.012 | 0.41 | 16.9 | 69 | 28 |
| Rhodothermia | class | 0.012 | 0.41 | 16.9 | 69 | 28 |
| Rhodothermales | order | 0.012 | 0.41 | 16.9 | 69 | 28 |
| Clostridium thermosuccinogenes | species | 0.008 | 0.37 | 16.3 | 57 | 21 |
| Pseudoclostridium | genus | 0.008 | 0.37 | 16.3 | 57 | 21 |
| Anaerovibrio | genus | 0.028 | 0.41 | 15.1 | 63 | 26 |
| Enterobacteriaceae | family | 0.055 | 2.12 | 12.7 | 32 | 68 |
| Enterobacterales | order | 0.057 | 2.12 | 12.7 | 32 | 68 |
| Peptoniphilus | genus | 0.052 | 0.46 | 12.6 | 65 | 30 |
| Porphyromonas canis | species | 0.005 | 0.39 | 12.1 | 46 | 18 |
| Bifidobacterium adolescentis | species | 0.011 | 2.18 | 12 | 28 | 61 |
| Phocaeicola massiliensis | species | 0.015 | 0.42 | 12 | 52 | 22 |
| Olivibacter | genus | 0.006 | 0.42 | 11.3 | 48 | 20 |
| Phocaeicola paurosaccharolyticus | species | 0.044 | 0.48 | 11.2 | 64 | 31 |
| Eukaryota | superkingdom | 0.002 | 3.5 | 11 | 8 | 28 |
Comparing Averages
A classic question on using averages: Do you include samples where a bacteria was not found as a zero, or exclude it from your average? I am inclined to suggest that both should be done.
| tax_name | Rank | Symptom Average | Reference Average | Symptom Average With Zero | Reference Average With Zero |
| Bifidobacterium catenulatum | species | 0.031 | 0.016 | 0.02 | 0.006 |
| Bifidobacterium gallicum | species | 0.845 | 0.293 | 0.695 | 0.142 |
| Bifidobacterium angulatum | species | 0.034 | 0.015 | 0.015 | 0.002 |
| Bifidobacterium asteroides | species | 0.007 | 0.005 | 0.003 | 0.001 |
| Caloramator indicus | species | 0.007 | 0.037 | 0.002 | 0.013 |
| Bifidobacterium cuniculi | species | 0.01 | 0.006 | 0.003 | 0.001 |
| Bifidobacterium subtile | species | 0.018 | 0.006 | 0.008 | 0.002 |
| Bacteroides rodentium | species | 0.187 | 0.397 | 0.182 | 0.366 |
| Phocaeicola paurosaccharolyticus | species | 0.035 | 0.06 | 0.033 | 0.055 |
| Bifidobacterium indicum | species | 0.028 | 0.019 | 0.021 | 0.009 |
| Hathewaya histolytica | species | 0.18 | 0.281 | 0.176 | 0.261 |
| Bifidobacterium scardovii | species | 0.005 | 0.009 | 0.002 | 0.001 |
| Leyella stercorea | species | 0.712 | 0.608 | 0.303 | 0.252 |
| Phocaeicola sartorii | species | 0.041 | 0.084 | 0.038 | 0.076 |
| Anaerovibrio lipolyticus | species | 0.055 | 0.114 | 0.047 | 0.098 |
| Phascolarctobacterium succinatutens | species | 0.033 | 0.061 | 0.022 | 0.045 |
| Sarcina maxima | species | 0.109 | 0.034 | 0.074 | 0.017 |
| Clostridium thermosuccinogenes | species | 0.008 | 0.015 | 0.006 | 0.012 |
| Butyricimonas virosa | species | 0.013 | 0.014 | 0.004 | 0.007 |
| Bifidobacterium adolescentis | species | 0.404 | 0.299 | 0.356 | 0.229 |
| Veillonella montpellierensis | species | 0.037 | 0.03 | 0.022 | 0.017 |
| Sporolactobacillus putidus | species | 0.039 | 0.018 | 0.015 | 0.005 |
| Caloramator uzoniensis | species | 0.005 | 0.01 | 0.002 | 0.005 |
| Johnsonella ignava | species | 0.065 | 0.051 | 0.063 | 0.047 |
| Bacteroides cellulosilyticus | species | 0.631 | 0.86 | 0.569 | 0.762 |
Averages to Median
Bacteria tend to have very high skewness rendering the use of means very unsafe. IMHO, medians is a far better statistics to use. As shown below, medians are almost always below the average because of extreme values. Often a mean will be at the 90%ile for a bacteria.
| tax_name | Rank | Symptom Avarage | Reference Average | Symptom Median | Reference Median |
| Bifidobacterium catenulatum | species | 0.031 | 0.016 | 0.01 | 0.003 |
| Bifidobacterium gallicum | species | 0.845 | 0.293 | 0.084 | 0.009 |
| Bifidobacterium angulatum | species | 0.034 | 0.015 | 0.009 | 0.004 |
| Bifidobacterium asteroides | species | 0.007 | 0.005 | 0.005 | 0.002 |
| Caloramator indicus | species | 0.007 | 0.037 | 0.002 | 0.005 |
| Bifidobacterium cuniculi | species | 0.01 | 0.006 | 0.006 | 0.003 |
| Bifidobacterium subtile | species | 0.018 | 0.006 | 0.007 | 0.003 |
| Bacteroides rodentium | species | 0.187 | 0.397 | 0.07 | 0.191 |
| Phocaeicola paurosaccharolyticus | species | 0.035 | 0.06 | 0.025 | 0.044 |
| Bifidobacterium indicum | species | 0.028 | 0.019 | 0.011 | 0.005 |
| Hathewaya histolytica | species | 0.18 | 0.281 | 0.092 | 0.159 |
| Bifidobacterium scardovii | species | 0.005 | 0.009 | 0.003 | 0.002 |
| Leyella stercorea | species | 0.712 | 0.608 | 0.005 | 0.003 |
| Phocaeicola sartorii | species | 0.041 | 0.084 | 0.017 | 0.033 |
| Anaerovibrio lipolyticus | species | 0.055 | 0.114 | 0.014 | 0.028 |
| Phascolarctobacterium succinatutens | species | 0.033 | 0.061 | 0.004 | 0.009 |
| Sarcina maxima | species | 0.109 | 0.034 | 0.017 | 0.007 |
| Clostridium thermosuccinogenes | species | 0.008 | 0.015 | 0.005 | 0.008 |
| Butyricimonas virosa | species | 0.013 | 0.014 | 0.011 | 0.006 |
| Bifidobacterium adolescentis | species | 0.404 | 0.299 | 0.044 | 0.011 |
| Veillonella montpellierensis | species | 0.037 | 0.03 | 0.019 | 0.007 |
| Sporolactobacillus putidus | species | 0.039 | 0.018 | 0.015 | 0.005 |
| Caloramator uzoniensis | species | 0.005 | 0.01 | 0.002 | 0.004 |
| Johnsonella ignava | species | 0.065 | 0.051 | 0.019 | 0.03 |
| Bacteroides cellulosilyticus | species | 0.631 | 0.86 | 0.02 | 0.079 |
Bottom Line
The purpose of this post is to discourage people from under-using the available data. Patterns can be counterintuitive, for example: Bifidobacteriums is detected less often, but when detected the amount is higher.
My goal for doing this deep dive is to look at different Odds Ratios. Odds ratios allows accurate prediction of likely and developing conditions. A sweet side effect is that it also allows priories on each bacteria to be objectively computed with a goal of higher success with interventions.
I have more investigations to do, especially double checking computations and doing cross validation with existing samples.
Recent Comments