This is part of a series of technical notes. If this interests you, you may wish to view others in the series.
A colleague wrote me:
I am currently researching companies who are using machine learning and LLM approaches for Microbiome analysis I wanted to know your opinion why this approach fails/ is less precise than your [MicrobiomePrescription MP] Fuzzy Logic Expert System with Monte Carlo method.
I have been doing a wide variety of Artificial Intelligence development professionally since 1988 for firms including Microsoft, Amazon, Verizon and Starbucks. I also have a reasonable science background including High School General Science teacher and College Chemistry and Physics instructor.
In the early days of Microbiome Prescription I tried a panacea of technics such as Random Forest, linear and non linear regression, supervised learning, etc. The results were less than acceptable. A friend, Richard Sprague, also ex-Microsoft, who worked as Citizen Scientist in Residence for uBiome for a while set up a series of meeting with the teams of the Allen Institute for AI [AI2]. The consensus working with those experts was that my direction of using the expert system model was far superior than what they could come up with.
How does Machine Learning and LLM work?
The core mechanism is pattern recognition of key words and phrases. This allows a numeric representation of the literature to be created, i.e. [subject #,verb #, object #]. When someone asks about subject, a set of equivalent subject # is obtained, a set of equivalent verb # is obtained and we just lookup the data. The number of records may be used to determine priority or most probable outcome.
This leads to the problem with many implementations, phrase recognition. To illustrate consider two articles, one mentions: Limosilactobacillus reuteri and the other mentions for Lactobacillus reuteri. A microbiologist knows that these are the same, but the typical ML or LLM does not. Do we end up with two collections of information? Since both Limosilactobacillus and Lactobacillus are also taxological units, do we get this species information incorrectly applied to the genus?
This is further complicated going to studies dealing with strains. Compare these two species: Escherichia coli O157:H7 and Escherichia coli Nissle 1917. One causes food poisoning, and the other is a probiotic shown to improves Crohn’s disease. Strains often have multiple identifiers and retail names. For example, Enterococcus faecium SF68 is also known as Enterococcus faecium NCIMB 10415 and sold retail as Bioflorin for humans and multiple brands in veterinary practice.
So the number 1 issue is correct identification. We also have some name collisions: Bacteroidetes is the name of both a class [Taxon 200643] and a phylum [Taxon 976] (which is now called Bacteroidota). A knowledgeable reader can reason out what is meant.
Taxonomy Hierarchy Inference
Information on the microbiome is sparse. A microbiome practitioner asked about what would reduce Lactobacillus balticus may discover that there is no literature on it. This practitioner would then infer that whatever reduces Lactobacillus would reduce it. This is an inference which MP does (remember it is an expert system mimicking the behavior of a human expert). MP takes it one step further by recognizing that is now classified as Limosilactobacillus balticus. Instead of items impacting Lactobacillus, it will use items impacting Limosilactobacillus. The suggestions are more probable to be correct.
What to address question
The various AI systems may well scrap ranges from studies to apply to a microbiome sample. Results are not consistent from lab to lab [the back story is this 2019 post: The taxonomy nightmare before Christmas…] . This means that reference ranges are more inconsistent as a consequence. Below are some range examples from commercial tests
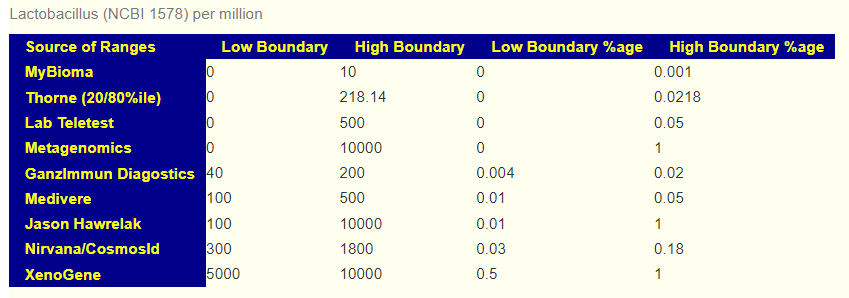
The reference range dilemma is why MP generates suggestions based on reference ranges determine by multiple methods: Average +/- Std Dev, Box Plot Whiskers, ranges from specific sources (including the processing lab in some cases), and patent pending algorithms. Each reference range is determined from a large collection of sample processed by the same lab. The suggestions using each of these reference ranges are then aggregated into a “consensus” report (i.e. Monte Carlo method).
“We are using AI to get Suggestions” – marketing hype!
A few years ago, microbiome testing companies would attempt to get creditability by claiming their suggestions were created by registered dietitian. Today, “AI” is the replacement. If you ask about what AI methodology is being used, the size and scope of the data behind, etc. 99% of the time you will be given a “It’s proprietary! We cannot disclose it“.
They may be truthful that is coming from AI, for example, someone asked https://www.perplexity.ai/, “Which foods reduces Fusobacterium” (Example answer – Fusobacterium nucleatum is what is cited). They copy the answer into their database to show their customers. Thus it is true that the answer is coming from AI; but it is a one-dimensional blinkered answer that will often leaves their customer worse. This AI is ignorant that Fusobacterium prausnitzii [The bacteria formerly known as Faecalibacterium prausnitzii] belongs to it. This bacteria has lots of studies that are ignored by the AI! The AI appears ignorant that two studies report both quercetin, mastic gum (prebiotic) reduces it. It likely has the data but has misclassified it.

My observations of reviewing many sites is that suggestions are scoped to a single bacteria and ignores side-effects on other bacteria that would also be impacted. MP uses holistic algorithms [which is what would be expected when someone has done Ph.D. courses on Integer and Non-Integer Programming Optimization]. The typical MP matrix to solve is around 60 taxa (up to 430 in some cases) by 2092 possible modifiers – thus an array of some 12,000 to 800,000 elements to consider. The Monte Carlo method typically uses 5 runs resulting in 60,000 to 4,000,000 elements evaluated.
Examples of LLM gone bad
Most experts know that Mutaflor is Escherichia coli Nissle 1917 and is clearly names as such in publicly accessible papers on the National Library of Medicine. So this response is one of AI’s famous hallucinations; hallucinations are not possible from expert systems.

Bottom Line
Microbiome testing firms may correctly claim[in a legal sense] they are AI based. If they refuse to fully disclose the methodology being used (ideally on their site), then the safest assumption is that they got a summer intern to ask one of the LLM’s the questions and just copied the answers into the database. Without full disclosure, they simply cannot be trusted.
If you are considering using AI because some “hot shot evangelist or venture capitalist” is pushing for it; then — look at the above issues and insists on documentation on how each of these issues will be addressed. Until there is clear, understandable documentation on these issues, “The suitability of AI has not been shown for the proposed AI implementation” and stop wasting time and money!
MP uses a very old model of AI that requires manual data curation being feed to the expert system. This a hallmark of expert systems.
Recent Comments